Beyond the Molecule: How BERT is Revolutionizing Materials and Drug Property Prediction
This article explores the transformative application of BERT-based architectures in predicting materials and molecular properties, a critical task in drug development and materials science.
Beyond the Molecule: How BERT is Revolutionizing Materials and Drug Property Prediction
Abstract
This article explores the transformative application of BERT-based architectures in predicting materials and molecular properties, a critical task in drug development and materials science. We first establish the foundational principles of adapting transformer models from natural language to chemical representations like SMILES. The discussion then progresses to methodological implementations, including multitask learning and cross-modal knowledge transfer, which address the pervasive challenge of data scarcity. Further, we delve into optimization strategies such as advanced positional embeddings and active learning integration to enhance model robustness and data efficiency. Finally, the article provides a comprehensive validation of BERT's performance against state-of-the-art graph neural networks and other traditional methods across diverse benchmarks, highlighting its superior accuracy and interpretability. This guide is tailored for researchers and professionals seeking to leverage cutting-edge deep learning for accelerated discovery.
From Text to Toxicity: The Foundational Shift to BERT in Materials Informatics
The Data Scarcity Problem in Drug and Materials Discovery
In both drug and materials discovery, researchers face a fundamental constraint: the scarcity of high-quality, labeled experimental data. This data scarcity problem creates a significant bottleneck in the development cycle, traditionally requiring years of laboratory experimentation and enormous financial investment to generate sufficient data for reliable predictive modeling. In drug discovery, the issue manifests in limited toxicology labels and clinical trial outcomes, while materials science grapples with sparse measurements of complex properties across vast chemical spaces. The high cost and extended timelines associated with experimental data generation—often requiring $10-100 million and 10-20 years to bring a new material to market—make this scarcity a critical barrier to innovation [1] [2].
Within this challenging landscape, BERT (Bidirectional Encoder Representations from Transformers) architecture and its derivatives have emerged as powerful frameworks for addressing data scarcity through representation learning and transfer learning. These models, initially pretrained on large unlabeled datasets, learn fundamental chemical and structural patterns that can be fine-tuned for specific prediction tasks with limited labeled examples. This approach has demonstrated remarkable success in both domains, effectively decoupling representation learning from downstream task-specific fine-tuning to overcome data limitations [3] [4] [5]. This article provides a comprehensive comparison of BERT-based approaches tackling the data scarcity problem, examining their experimental methodologies, performance benchmarks, and practical implementations.
Comparative Analysis of BERT-Based Solutions
Table 1: Overview of BERT-Based Architectures for Property Prediction
| Model Name | Architectural Features | Pretraining Data | Target Applications | Key Innovations |
|---|---|---|---|---|
| Molecular BERT [3] | Transformer-based BERT | 1.26 million compounds | Drug toxicity prediction | Disentangles representation learning and uncertainty estimation |
| GEO-BERT [4] | Geometry-enhanced BERT | Molecular structures with 3D conformations | Drug discovery (DYRK1A inhibitors) | Incorporates atom-atom, bond-bond, and atom-bond positional relationships |
| Cross-modal BERT [5] | Multimodal BERT with knowledge transfer | Multimodal materials data | Composition-based materials property prediction | Aligns compositional and structural embeddings through implicit/explicit transfer |
| CrystalTransformer [6] | Transformer-generated atomic embeddings | Crystal structures from materials databases | Crystal property prediction | Generates universal atomic embeddings (ct-UAEs) transferable across properties |
Table 2: Experimental Performance Benchmarks of BERT Models
| Model | Dataset | Key Metrics | Performance Improvement | Data Efficiency Advantage |
|---|---|---|---|---|
| Molecular BERT [3] | Tox21, ClinTox | Toxic compound identification | Achieved equivalent performance with 50% fewer iterations vs conventional AL | Reliable uncertainty estimation with limited labeled data |
| GEO-BERT [4] | DYRK1A inhibitor screening | IC50 values (<1 μM) | Identified two potent novel inhibitors in prospective validation | Enhanced molecular characterization from 3D structural information |
| Cross-modal BERT [5] | LLM4Mat-Bench (32 tasks) | Mean Absolute Error (MAE) | State-of-the-art in 25 out of 32 cases, MAE reduced by 15.7% on average | Effective knowledge transfer from compositional to structural domains |
| CrystalTransformer [6] | Materials Project database | Formation energy prediction | 14% improvement in CGCNN, 18% in ALIGNN with ct-UAEs | Addresses data scarcity through transferable atomic fingerprints |
Experimental Protocols and Methodologies
Molecular BERT for Active Learning in Drug Discovery
The Molecular BERT framework employs a sophisticated Bayesian experimental design integrated with active learning to address data scarcity in toxicity prediction [3]. The methodology begins with pretraining a transformer-based BERT model on 1.26 million unlabeled compounds, enabling the model to learn fundamental chemical representations without labeled data. For downstream tasks, the implementation uses a small initial labeled set (100 molecules with balanced positive/negative instances) from Tox21 and ClinTox datasets, with the remaining training data forming an unlabeled pool set.
The experimental workflow applies scaffold splitting with an 80:20 ratio to create distinct training and testing sets, ensuring that molecules with similar core structures are segregated between sets to test generalization capability. The active learning cycle employs Bayesian acquisition functions to strategically select the most informative samples from the unlabeled pool:
- BALD (Bayesian Active Learning by Disagreement): Selects samples that maximize information gain about model parameters, calculated as
BALD(x) = H[y|x,D] - E_φ∼p(φ|D)H[y|x,φ]where the first term represents total uncertainty and the second term captures aleatoric uncertainty [3]. - EPIG (Expected Predictive Information Gain): Prioritizes samples expected to most improve predictive performance on target distributions by explicitly reducing model output uncertainty.
Through iterative cycles of sample selection, labeling, and model retraining, this approach achieves progressive improvement in predictive accuracy while minimizing labeling efforts. The disentanglement of representation learning (handled during pretraining) from uncertainty estimation (managed during active learning) enables reliable molecule selection despite limited initial labeled data [3].
GEO-BERT Geometry-Enhanced Molecular Representation
GEO-BERT addresses data scarcity by incorporating three-dimensional structural information through a self-supervised learning framework [4]. The model enhances its ability to characterize molecular structures by introducing three distinct positional relationships derived from 3D conformations:
- Atom-atom relationships: Spatial proximities and interactions between atoms
- Bond-bond relationships: Geometric arrangements between chemical bonds
- Atom-bond relationships: Relative orientations between atoms and bonds
The experimental validation involved prospective studies for DYRK1A inhibitor discovery, where the model was tasked with identifying novel inhibitors from chemical libraries. The methodology included transfer learning from the pretrained GEO-BERT to specific property prediction tasks with limited labeled examples, demonstrating that geometric pretraining provides robust molecular representations that transfer effectively to low-data scenarios. The model's open-source implementation (https://github.com/drug-designer/GEO-BERT) has proven practical utility in early-stage drug discovery, with experimental confirmation of two potent inhibitors (IC50: <1 μM) identified through this approach [4].
Cross-Modal Knowledge Transfer for Materials Property Prediction
For materials discovery where crystal structure data is often scarce, cross-modal BERT approaches address data scarcity through knowledge transfer between different representations of materials [5]. The methodology implements two distinct transfer learning strategies:
- Implicit Transfer (imKT): Involves pretraining chemical language models on multimodal embeddings aligned with a foundation model trained on multiple materials modalities (crystal structure, density of electronic states, charge density, and textual descriptions).
- Explicit Transfer (exKT): Generates crystal structures from composition using a large language model (CrystaLLM) as a crystal structure predictor, followed by structure-aware predictors (graph neural networks) fine-tuned on the generated crystals.
The experimental protocol evaluated these approaches on the LLM4Mat-Bench and MatBench datasets, encompassing 32 different prediction tasks. For composition-based property prediction, the models were trained using masked language modeling objectives on stoichiometric formulas, then fine-tuned for specific property prediction tasks. This approach demonstrated particularly strong performance on band gap-related predictions, with MAE reductions of 15.2% on average compared to previous state-of-the-art models [5].
CrystalTransformer for Universal Atomic Embeddings
CrystalTransformer addresses data scarcity in crystal property prediction through transferable atomic embeddings called universal atomic embeddings (ct-UAEs) [6]. The methodology involves:
- Pretraining: The CrystalTransformer model learns atomic embeddings directly from chemical information in crystal databases without relying on predefined atomic attributes.
- Transfer Learning: The generated ct-UAEs are transferred to various graph neural network backends (CGCNN, MEGNET, ALIGNN) for specific property prediction tasks.
- Multi-task Learning: Embeddings are trained across multiple properties to enhance transferability across different prediction tasks.
The experimental validation used Materials Project datasets (MP and MP) with standard splits (60,000 training, 5,000 validation, 4,239 testing for MP; 80%/10%/10% split for MP). Results demonstrated that ct-UAEs achieve significant accuracy improvements across multiple back-end models and properties, with the largest improvement (18% MAE reduction) observed in ALIGNN for formation energy prediction. The embeddings also showed excellent transferability across databases, with a 34% accuracy boost in MEGNET when applied to hybrid perovskites database [6].
The Scientist's Toolkit: Essential Research Reagents and Solutions
Table 3: Key Research Reagents and Computational Tools
| Reagent/Solution | Function | Application Context |
|---|---|---|
| Tox21 Dataset [3] | Provides ~8,000 compounds with 12 toxicity pathway measurements | Benchmark for computational toxicology models |
| ClinTox Dataset [3] | Contains 1,484 FDA-approved and failed clinical trial drugs | Drug safety profiling and toxicity prediction |
| Materials Project Database [6] | Computational repository with 134,243+ material structures and properties | Training and validation for materials property prediction |
| OMol25 Dataset [2] | Contains over 100 million DFT evaluations across ~83 million molecular systems | Training machine-learned interatomic potentials with near-DFT accuracy |
| Bayesian Active Learning Framework [3] | Strategically selects informative samples for labeling to minimize experimental costs | Active learning cycles for iterative model improvement |
| Universal Atomic Embeddings (ct-UAEs) [6] | Transferable atomic fingerprints capturing complex atomic features | Enhancing prediction accuracy across multiple GNN architectures |
| Cross-modal Alignment [5] | Bridges compositional and structural representations of materials | Property prediction for compounds without known crystal structures |
Technical Implementation and Workflow Integration
The technical implementation of BERT-based solutions for data scarcity follows a consistent pattern across drug and materials discovery domains. The fundamental approach involves decoupling representation learning from task-specific fine-tuning, which proves particularly valuable in low-data regimes. Implementation typically begins with self-supervised pretraining on large unlabeled datasets—1.26 million compounds for Molecular BERT or extensive crystal structure databases for CrystalTransformer—to learn fundamental chemical and structural patterns without expensive experimental labels [3] [6].
For specific property prediction tasks, the pretrained models undergo fine-tuning with limited labeled examples, leveraging the acquired representations to achieve robust performance with minimal task-specific data. In drug discovery applications, this process is often enhanced through Bayesian active learning frameworks that strategically select the most informative samples for experimental labeling, maximizing information gain while minimizing labeling costs [3]. The BALD and EPIG acquisition functions play crucial roles in this process, quantifying different aspects of uncertainty to guide sample selection.
In materials discovery, cross-modal knowledge transfer enables prediction for compounds without known structures by aligning compositional and structural representations [5]. The implicit transfer approach (imKT) aligns chemical language model embeddings with multimodal foundation models, while explicit transfer (exKT) generates plausible crystal structures from composition alone. This enables structure-aware property prediction even when experimental structure determinations are unavailable, significantly expanding the explorable chemical space.
BERT-based architectures have fundamentally transformed the approach to data scarcity in drug and materials discovery, demonstrating that representation learning and transfer learning can effectively mitigate the challenges of limited experimental data. The comparative analysis reveals that while architectural variants differ in their specific implementations—incorporating geometric information, cross-modal transfer, or universal embeddings—they share a common foundation of pretraining followed by task-specific adaptation.
The most successful approaches effectively disentangle representation learning from uncertainty estimation, enabling robust performance even with limited labeled examples. Molecular BERT's 50% reduction in required iterations for equivalent toxicity identification, GEO-BERT's experimental validation through novel inhibitor discovery, and CrystalTransformer's 14-18% accuracy improvements across multiple graph neural network architectures collectively demonstrate the transformative potential of these approaches [3] [4] [6].
As these technologies evolve, key challenges remain in improving interpretability, enhancing multimodal integration, and developing more sophisticated uncertainty quantification methods. However, the current state of BERT-based property prediction already offers powerful solutions to the data scarcity problem, enabling more efficient exploration of chemical and materials spaces while significantly reducing the experimental burden required for discovery and development.
In the realm of computational chemistry and drug discovery, the Simplified Molecular Input Line Entry System (SMILES) has established itself as a fundamental vocabulary for representing molecular structures. Much like natural language processing (NLP) models operate on sequences of words, chemical language models (CLMs) utilize SMILES strings as their foundational linguistic elements. These strings encode two-dimensional molecular information through a specialized vocabulary of characters ("tokens") that represent atoms, bonds, rings, and branches [7]. The SMILES notation functions as a specialized chemical grammar, with specific syntax rules governing how tokens can be combined to form valid molecular representations. This linguistic analogy extends to how researchers can apply NLP-inspired techniques—including data augmentation, token manipulation, and semantic analysis—to enhance model performance in critical tasks such as materials property prediction and drug-target interaction (DTI) forecasting [7] [8].
Within BERT-based architectures for materials property prediction, understanding SMILES as a vocabulary is not merely an abstract concept but a practical framework that drives methodological innovation. The representation of molecules as sequences enables the application of transformer-based models that can capture complex, long-range dependencies within molecular structures [9]. This approach has demonstrated significant potential in addressing one of the field's most pressing challenges: achieving accurate predictions with limited labeled data. By leveraging pre-trained chemical language models, researchers can transfer knowledge from large unlabeled molecular datasets to specific property prediction tasks, substantially improving data efficiency and model generalization [9].
The SMILES Lexicon: Tokenization and Vocabulary Construction
Fundamental Token Types
The SMILES vocabulary consists of distinct token types that collectively describe molecular structure:
- Element tokens: Represent atomic species (e.g., "C" for carbon, "N" for nitrogen, "O" for oxygen), with aromaticity distinguished by lowercase characters ("c" for aromatic carbon) [10].
- Bond tokens: Describe connection types ("-" for single bonds, "=" for double bonds, "#" for triple bonds) with single bonds often omitted as defaults [7].
- Branching tokens: Parentheses "(" and ")" indicate molecular branching patterns [7].
- Ring tokens: Numeric labels (e.g., "1", "2") mark ring closure points within the structure [7].
This grammatical framework allows SMILES to represent complex molecular graphs as linear strings through depth-first traversal of the molecular structure [7]. A single molecule can generate multiple valid SMILES strings depending on the starting atom and traversal path, creating inherent synonymity within the chemical language [7].
Advanced Tokenization Strategies
Recent research has evolved beyond basic SMILES tokenization to address vocabulary limitations. The Atom-In-SMILES (AIS) approach enhances token informativeness by incorporating local chemical environment context into each token [10]. Unlike standard SMILES tokens that represent atoms in isolation, AIS tokens encapsulate three key aspects: the elemental symbol of the central atom, ring membership information ("R" for ring atoms, "!R" for non-ring atoms), and the neighboring atoms connected to the central atom [10]. This environment-aware tokenization creates a more chemically meaningful vocabulary while maintaining SMILES grammar compatibility.
Hybrid representation methods such as SMI+AIS(N) selectively replace frequently occurring SMILES tokens with their AIS counterparts, balancing chemical expressiveness with vocabulary size [10]. This approach mitigates the significant token frequency imbalance inherent in standard SMILES, where common tokens like "C" (carbon) appear with disproportionately high frequency compared to other elements [10].
Table 1: Comparison of Molecular Representation Methods
| Representation | Token Diversity | Chemical Context | Validity Guarantee | Primary Applications |
|---|---|---|---|---|
| Standard SMILES | Limited | Minimal | No | General molecular representation |
| SELFIES | Limited | Minimal | Yes | Robust molecular generation |
| AIS | High | Extensive | No | Property prediction tasks |
| SMI+AIS | Moderate | Selective | No | Structure generation optimization |
Quantitative Performance Comparison of SMILES Representation Methods
Structure Generation and Optimization Performance
The effectiveness of different SMILES representations has been quantitatively evaluated in molecular structure generation tasks. When applied to latent space optimization with Bayesian optimization for generating structures with improved binding affinity and synthesizability, the SMI+AIS representation demonstrated measurable advantages over established alternatives [10]. Specifically, SMI+AIS achieved a 7% improvement in binding affinity and a 6% increase in synthesizability scores compared to standard SMILES representations [10]. This performance enhancement stems from the richer chemical context encoded within AIS tokens, which allows optimization algorithms to better capture structure-property relationships.
The hybridization approach in SMI+AIS also addresses vocabulary imbalance issues that can impede model training. Analysis of the ZINC database revealed that introducing 100-150 carefully selected AIS tokens effectively redistributes token frequencies, creating a more balanced vocabulary without excessive expansion that could lead to data sparsity issues [10]. This balanced vocabulary composition correlates with improved model performance in downstream tasks.
Data Augmentation Strategies and Their Effects
SMILES enumeration (generating multiple valid representations of the same molecule) has emerged as a powerful data augmentation technique, particularly beneficial in low-data scenarios [7]. Beyond simple enumeration, researchers have developed sophisticated augmentation strategies that further enhance model performance:
Table 2: Performance of SMILES Augmentation Strategies in Low-Data Scenarios
| Augmentation Method | Validity | Uniqueness | Novelty | Optimal Probability (p) |
|---|---|---|---|---|
| Token Deletion | Variable | High | High | 0.05 |
| Atom Masking | High | High | Moderate | 0.05 |
| Bioisosteric Substitution | High | Moderate | Moderate | 0.15 |
| Self-training | Highest | High | High | N/A |
These augmentation strategies exhibit distinct performance characteristics across dataset sizes. Atom masking has proven particularly effective for learning desirable physicochemical properties in very low-data regimes, while token deletion shows promise for creating novel molecular scaffolds [7]. Self-training augmentation, wherein SMILES strings generated by a chemical language model are used as input for subsequent training phases, consistently outperforms basic enumeration across all dataset sizes [7].
Experimental Protocols for SMILES-Based Molecular Representation
SMILES Alignment Protocol for Molecular Similarity
A key methodology for comparing SMILES-represented molecules adapts the Needleman-Wunsch algorithm for global sequence alignment with a modified scoring function [11]. This approach enables quantitative assessment of molecular transformations in biochemical pathways:
- Input Preparation: Convert molecules to canonical SMILES representations using standard toolkits.
- Scoring Matrix Definition: Implement a substitution matrix based on atomic partial charges rather than simple atomic identities, reflecting electronegativity differences.
- Gap Penalty Configuration: Assign appropriate gap opening and extension penalties to balance alignment flexibility with chemical plausibility.
- Dynamic Programming Execution: Perform global alignment using the modified scoring scheme.
- Pathway Analysis Application: Quantify structural changes between metabolites in pathways such as glycolysis and the Krebs cycle.
This method has validated its efficacy by correctly aligning atoms known to be conserved across biochemical transformations, successfully capturing the structural evolution patterns characteristic of linear versus cyclical metabolic pathways [11].
Chemical Language Model Pretraining and Fine-Tuning
Effective implementation of BERT-style architectures for molecular property prediction follows a rigorous protocol:
- Large-Scale Pretraining: Train transformer models on extensive unlabeled molecular datasets (e.g., 1.26 million compounds for MolBERT) using masked language modeling objectives [9].
- Embedding Alignment: For materials property prediction, align CLM embeddings with those from multimodal foundation models incorporating crystal structure, electronic states, and textual descriptions [5].
- Task-Specific Fine-Tuning: Adapt pretrained models to specific property prediction tasks using limited labeled data, typically with scaffold-based dataset splits to ensure generalization [9].
- Bayesian Active Learning Integration: Employ acquisition functions like Bayesian Active Learning by Disagreement (BALD) to strategically select informative molecules for labeling, maximizing model improvement per experiment [9].
This protocol has demonstrated remarkable data efficiency, achieving equivalent toxic compound identification with 50% fewer iterations compared to conventional active learning approaches [9].
SMILES Processing in Chemical Language Models
SMILES-Enhanced Architectures for Property Prediction
Cross-Modal Knowledge Transfer Frameworks
Advanced SMILES-based prediction systems increasingly employ cross-modal knowledge transfer to enhance performance. Two predominant formulations have emerged:
- Implicit Transfer (imKT): Involves pretraining chemical language models on multimodal embeddings aligned with foundation models incorporating multiple materials modalities (crystal structure, density of electronic states, charge density, textual descriptions) [5].
- Explicit Transfer (exKT): Generates crystal structures using large language models like CrystaLLM, followed by structure-aware predictor fine-tuning on generated crystals [5].
These approaches have demonstrated state-of-the-art performance on benchmark datasets, achieving mean absolute error reductions of 15.7% on JARVIS-DFT tasks and 15.2% on SNUMAT band-gap prediction tasks compared to previous benchmarks [5]. The integration of SMILES representations with multimodal knowledge creates more robust and accurate property prediction systems.
Hybrid Model Architectures for Specialized Applications
Sophisticated hybrid architectures have emerged for specific drug discovery applications:
SVDTI Framework: This drug-target interaction prediction model employs a stacked variational autoencoder (SVAE) with Long Short-Term Memory (LSTM) networks to map high-dimensional SMILES and protein sequence data into compact, informative low-dimensional vectors [12]. The framework subsequently processes these representations through a neural collaborative filtering (NCF) model that combines the linear characteristics of matrix factorization with the nonlinear representation power of multilayer perceptrons [12].
Imagand Model: This SMILES-to-Pharmacokinetic (S2PK) diffusion model generates pharmacokinetic properties conditioned on learned SMILES embeddings, addressing the challenge of sparse PK datasets with limited overlap [13]. The model employs a Discrete Local Gaussian Noise (DLGN) approach that creates a prior distribution closer to the true data distribution, improving generation performance for non-Gaussian distributed molecular properties [13].
SVDTI Framework for Drug-Target Interaction Prediction
Table 3: Key Research Resources for SMILES-Based Molecular Modeling
| Resource | Type | Function | Application Context |
|---|---|---|---|
| RDKit | Software Library | Molecular fingerprint generation & manipulation | Similarity analysis, descriptor calculation [14] |
| Yamanishi Dataset | Curated Dataset | Gold-standard drug-target interactions | Model benchmarking & validation [12] |
| ZINC Database | Molecular Database | Large collection of commercially available compounds | Vocabulary analysis & model pretraining [10] |
| SwissBioisostere | Specialized Database | Bioisosteric replacement patterns | Data augmentation strategy [7] |
| Tox21/ClinTox | Benchmark Datasets | Toxicology & clinical failure data | Model evaluation & validation [9] |
| MolBERT | Pretrained Model | Chemical language model with 1.26M compounds | Transfer learning initialization [9] |
The evolving understanding of SMILES as a specialized vocabulary continues to drive innovation in chemical language modeling. Current research demonstrates that moving beyond basic tokenization toward environmentally aware representations like AIS tokens and hybrid SMI+AIS approaches yields measurable performance improvements in critical tasks including molecular generation, property prediction, and drug-target interaction forecasting [10]. The integration of SMILES processing with multimodal knowledge transfer and sophisticated architectures like stacked variational autoencoders and diffusion models represents the cutting edge of computational molecular design [5] [12] [13].
As the field advances, the SMILES vocabulary is likely to further evolve toward increasingly context-aware representations that capture richer chemical semantics while maintaining compatibility with the extensive existing ecosystem of computational tools. These developments will strengthen the foundation for more accurate, data-efficient, and interpretable molecular property prediction systems, ultimately accelerating the drug and materials discovery pipeline.
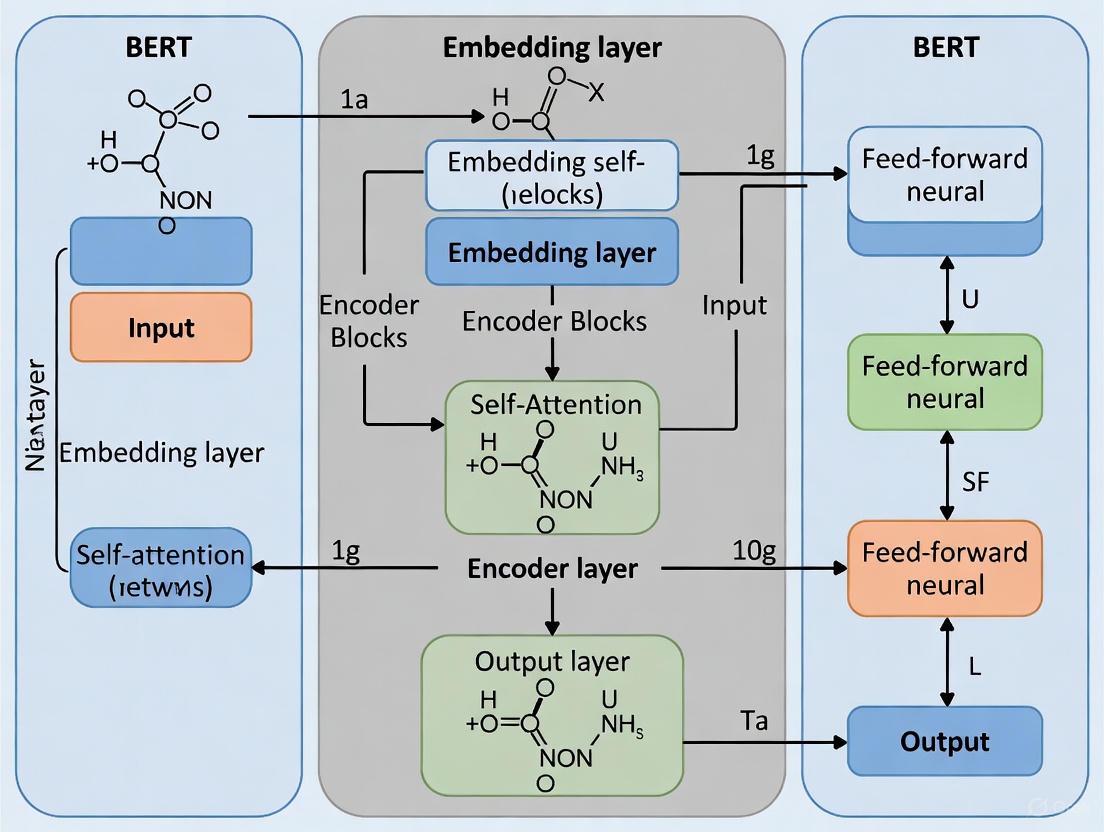
The Bidirectional Encoder Representations from Transformers (BERT) represents a fundamental shift in how machines understand human language. Introduced by Google in 2018, its core innovation lies in its bidirectional context processing and sophisticated self-attention mechanism [15]. Unlike previous models that processed text sequentially (either left-to-right or right-to-left), BERT's key innovation is its ability to read an entire sequence of words at once [15]. This non-directional approach enables the model to learn a deeper context of a word by considering all of its surroundings simultaneously [15].
In the specific context of materials property prediction and drug discovery, this architectural advantage translates into a powerful ability to understand complex molecular representations and clinical text data. Models like GEO-BERT and DrugBERT, built upon the core BERT architecture, leverage these capabilities to predict molecular properties and drug efficacy with remarkable accuracy [4] [16]. This guide will objectively compare BERT's performance against alternative architectures and provide detailed experimental protocols from recent research, focusing specifically on applications in scientific property prediction.
Core Architectural Components
The Self-Attention Mechanism
At the heart of BERT lies the self-attention mechanism, which allows the model to weigh the importance of different words in a sequence when encoding a particular word [17]. In technical terms, self-attention is a mechanism where each token in the input pays attention to all other tokens, including itself, to generate its contextual embedding [17]. Calculating attention is a way for each token to ask, "Which other words should I focus on to understand my meaning?"
The mechanism operates through three learned vectors for each token:
- Query (Q): Represents what the current token is looking for in other tokens
- Key (K): Helps other tokens decide how relevant the current token is to them
- Value (V): Contains the actual information that a token contributes [17]
These vectors are computed using learned weight matrices (Wq, Wk, W_v) during training. The attention score is calculated by taking the dot product of the query vector of one token with the key vector of another, then applying a softmax function to obtain normalized weights [18].
Bidirectional Context Processing
BERT's bidirectionality is fundamentally different from the unidirectional approach of models like GPT. While GPT processes text strictly from left to right, BERT's encoder-only architecture processes all words in a sequence simultaneously [15]. This bidirectional training enables BERT to develop a deeper understanding of language context, making it particularly effective for tasks that require comprehensive contextual analysis rather than text generation [15].
The bidirectional capability is achieved through BERT's pre-training tasks:
- Masked Language Modeling (MLM): Approximately 15% of words in input sequences are randomly masked, and the model must predict these masked words based on their bidirectional context [15]
- Next Sentence Prediction (NSP): The model receives pairs of sentences and predicts whether the second sentence logically follows the first [15]
Comparative Performance Analysis
Benchmark Performance Across Domains
Extensive testing across multiple domains reveals distinct performance patterns for BERT and its alternatives. The following table summarizes key comparative findings:
Table 1: Performance comparison of BERT and alternative models across different domains and tasks
| Model | Architecture Type | Primary Strengths | Notable Performance Metrics | Domain Applications |
|---|---|---|---|---|
| BERT | Encoder-only, Bidirectional | Deep contextual understanding, NLU tasks | Superior performance on medical concept recognition vs. general BERT [19] | Drug discovery, molecular property prediction [4] |
| GEO-BERT | BERT-based with geometric encoding | Molecular property prediction, 3D structure integration | Identified potent DYRK1A inhibitors (IC50: <1 μM) [4] | Drug discovery, molecular analysis [4] |
| GPT Series | Decoder-only, Unidirectional | Text generation, creative tasks | 87% accuracy in clinical sentiment classification [20] | Content creation, conversational AI [15] |
| LLaMA | Decoder-only, Autoregressive | Computational efficiency, strong performance with fewer parameters | Comparable performance to larger models with fewer parameters [15] | Accessible AI research, resource-constrained environments [15] |
| BioBERT | Domain-specific BERT | Biomedical text processing | F1-score of 0.836 on clinical trial NER [21] | Clinical text analysis, biomedical NER [21] |
| DrugBERT | BERT with LDA topic embedding | Drug efficacy prediction | 3% improvement in AUC over previous methods [16] | Anti-tumor drug efficacy prediction [16] |
Domain-Specific Performance in Scientific Applications
In specialized scientific domains, BERT-based models consistently demonstrate advantages over general-purpose alternatives:
Table 2: Performance of BERT-based models in specialized scientific applications
| Application Domain | Model Variant | Task | Performance Metrics | Comparative Advantage |
|---|---|---|---|---|
| Molecular Property Prediction | GEO-BERT [4] | Molecular property prediction, inhibitor identification | Identified two novel DYRK1A inhibitors with IC50 <1 μM [4] | Incorporates 3D structural information via atom-atom, bond-bond, and atom-bond relationships [4] |
| Drug Efficacy Prediction | DrugBERT [16] | Predicting efficacy of anti-tumor drugs | 3% AUC improvement on independent bowel cancer dataset [16] | Integrates LDA topic embedding and drug efficacy-aware attention mechanism [16] |
| Clinical Text Analysis | BioBERT/ClinicalBERT [19] | Medical concept recognition | Outperformed general BERT; ClinicalBERT achieved mean macro-F1 score of 0.761 [19] | Domain-specific pre-training on biomedical corpora [19] |
| Clinical Trial NER | PubMedBERT [21] | Named Entity Recognition in eligibility criteria | F1-scores of 0.715, 0.836, and 0.622 across three corpora [21] | Superior to both general BERT and other biomedical variants [21] |
Experimental Protocols and Methodologies
GEO-BERT for Molecular Property Prediction
The GEO-BERT framework exemplifies how core BERT architecture can be adapted for molecular property prediction in drug discovery. The experimental protocol involves several sophisticated components:
Molecular Representation: GEO-BERT considers atoms and chemical bonds in chemical structures as input, integrating positional information from three-dimensional molecular conformations [4]. Specifically, it introduces three different positional relationships: atom-atom, bond-bond, and atom-bond [4].
Architecture Enhancements:
- Self-supervised representation learning framework based on BERT
- Incorporation of 3D structural information within molecules
- Pre-training on large-scale small molecule data [4]
Experimental Validation:
- Benchmarking studies across multiple benchmarks demonstrated optimal performance
- Prospective validation through screening for DYRK1A inhibitors
- Discovery of two potent and novel DYRK1A inhibitors (IC50: <1 μM) confirmed practical utility [4]
DrugBERT for Anti-Tumor Drug Efficacy Prediction
DrugBERT represents another BERT-based adaptation specifically designed for predicting anti-tumor drug efficacy based on clinical text data:
Architecture Modifications:
- Integration of LDA-generated topic embeddings as semantic enhancement modules
- Drug efficacy-aware attention mechanism to prioritize drug efficacy-related semantic features
- LSTM integration to capture long-range dependencies in clinical text data [16]
Experimental Setup:
- Dataset: 958 patients with non-small cell cancer treated with anti-tumor drugs
- Independent validation: 266 bowel cancer patients
- Addressing data imbalance using SMOTE algorithm to synthesize minority class samples [16]
Methodological Innovation: The drug efficacy-aware attention mechanism enhances attention weights between drug efficacy relevant keywords. From K topics, m topics demonstrating significant drug efficacy relevance are selected, with the top w probability-ranked words extracted from chosen topics [16]. After deduplication, a Drug Efficacy-Related Keyword Repository (DEKR) containing n unique keywords is constructed [16].
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential research reagents and computational tools for BERT-based molecular property prediction
| Tool/Resource | Type | Primary Function | Application Example |
|---|---|---|---|
| GEO-BERT Framework [4] | Software Framework | Molecular property prediction with 3D structural integration | Predicting molecular properties and identifying DYRK1A inhibitors in early-stage drug discovery [4] |
| DrugBERT Framework [16] | Software Framework | Drug efficacy prediction from clinical text | Predicting efficacy of anti-tumor drugs based on clinical radiomic text data [16] |
| LDA Topic Model [16] | Computational Algorithm | Extracting latent topics from text corpora | Generating topic embeddings for semantic enhancement in DrugBERT [16] |
| SMOTE Algorithm [16] | Data Preprocessing | Addressing class imbalance in datasets | Synthesizing minority class samples in clinical trial data [16] |
| BioBERT/ClinicalBERT [19] [21] | Pre-trained Models | Domain-specific natural language processing | Medical concept recognition and named entity recognition in clinical text [19] [21] |
| SHAP (SHapley Additive exPlanations) [18] | Model Interpretation | Explaining model predictions based on game theory | Providing interpretability for BERT-based model predictions in academic assessment [18] |
The core BERT architecture, with its fundamental components of self-attention and bidirectional context processing, provides a powerful foundation for scientific property prediction research. The experimental evidence demonstrates that BERT-based models consistently outperform general-purpose alternatives in specialized domains such as molecular property prediction and drug efficacy assessment [4] [16].
The success of domain-specific adaptations like GEO-BERT and DrugBERT highlights the importance of architectural customization for scientific applications. By integrating domain knowledge through geometric representations [4] or topic-aware attention mechanisms [16], researchers can leverage BERT's core strengths while addressing specific challenges in materials science and drug development.
For research teams working in molecular property prediction, the evidence suggests that BERT-based architectures provide a robust foundation that can be productively specialized through domain-specific modifications. The bidirectional context understanding that defines BERT appears particularly valuable for analyzing complex molecular structures and clinical text data, making it an enduring architectural paradigm for scientific AI applications.
The Bidirectional Encoder Representations from Transformers (BERT) model, renowned for its revolutionary impact on natural language processing (NLP), is now pioneering a transformative shift in scientific computation, particularly in molecular property prediction for drug discovery. Originally designed for masked language modeling (MLM) tasks, BERT's core architecture possesses a unique capability to learn profound contextual relationships from sequential data. This intrinsic strength has enabled its successful adaptation from textual sequences to the structural "languages" of science—namely, the sequences of atoms and bonds that define chemical compounds. The adaptation of BERT for scientific applications represents a significant paradigm shift, moving beyond traditional quantitative structure-property relationship (QSPR) models that rely on hand-crafted descriptors towards deep learning approaches that learn optimal structure-to-descriptor mappings directly from data [9]. This guide provides a comprehensive comparison of emerging BERT-based frameworks for molecular property prediction, detailing their experimental performance, methodologies, and practical implementations to inform researchers and drug development professionals.
Understanding the Foundation: Masked Language Modeling
Masked language modeling serves as the foundational pre-training objective that enables BERT's sophisticated contextual understanding. In standard NLP applications, MLM involves randomly masking a portion of input tokens (typically 15%) and training the model to predict the original vocabulary identifiers of these masked tokens based on their bidirectional context [22] [23]. This self-supervised approach forces the model to develop a deep, bidirectional understanding of sequential relationships without requiring labeled datasets. The model achieves this by generating probability distributions over the input vocabulary for each masked token and minimizing the prediction error against the original tokens [22]. This pre-training paradigm has proven exceptionally transferable to molecular representations, where atoms or molecular fragments can be treated as "words" and entire molecular structures as "sentences," creating a powerful framework for learning complex chemical relationships from large unannotated molecular datasets [9].
Comparative Analysis of Molecular BERT Frameworks
Performance Benchmarking
Recent research has yielded several specialized BERT adaptations for molecular property prediction. The table below summarizes the key performance metrics of these frameworks across established benchmarks.
Table 1: Performance Comparison of BERT-based Molecular Property Prediction Models
| Model Name | Architectural Features | Benchmark Datasets | Key Performance Results | Computational Requirements |
|---|---|---|---|---|
| GEO-BERT [4] | Incorporates 3D molecular conformation data; Atom-atom, bond-bond, and atom-bond positional relationships | Multiple benchmarks (unspecified); DYRK1A inhibitor case study | "Optimal performance across multiple benchmarks"; Identified two novel DYRK1A inhibitors (IC50: <1 μM) | Requires 3D structural information |
| Pretrained BERT + Bayesian AL [9] [24] | BERT pretrained on 1.26M compounds combined with Bayesian active learning | Tox21; ClinTox | Achieved equivalent toxic compound identification with 50% fewer iterations vs. conventional active learning | Pretraining on large dataset; efficient fine-tuning |
| Ensemble Model (BERT, RoBERTa, XLNet) [25] | Ensemble learning with BERT, RoBERTa, and XLNet without extensive pretraining | Molecular property prediction tasks | "Significant effectiveness compared to existing advanced models"; addresses limited computational resources | Resource-efficient; no extensive pretraining needed |
Experimental Workflows and Methodologies
GEO-BERT Experimental Protocol
GEO-BERT introduces a geometry-aware framework that incorporates three-dimensional molecular conformation data into the BERT architecture [4]. The methodology involves:
- Molecular Representation: Atoms and chemical bonds in chemical structures serve as input, with integration of three-dimensional conformational positional information.
- Positional Relationships: Implementation of three novel positional encoding types - atom-atom, bond-bond, and atom-bond relationships - to enhance molecular structure characterization.
- Pre-training Strategy: Self-supervised pre-training on large-scale small molecule datasets using the MLM objective, incorporating 3D structural information.
- Fine-tuning: Supervised fine-tuning on specific property prediction tasks, such as DYRK1A inhibitor identification.
The model's effectiveness was validated through prospective studies identifying novel DYRK1A inhibitors, with two compounds demonstrating potent inhibition (IC50: <1 μM) [4].
Pretrained BERT with Bayesian Active Learning Protocol
This approach integrates transformer-based BERT pretrained on 1.26 million compounds into a Bayesian active learning pipeline [9]:
Data Preparation:
- Utilizes Tox21 (≈8,000 compounds, 12 toxicity pathways) and ClinTox (1,484 compounds) datasets.
- Implements scaffold splitting with 80:20 ratio for training and testing to evaluate generalization.
- Constructs a balanced initial set of 100 molecules with equal positive/negative representation.
Model Architecture:
- Employs MolBERT, a BERT adaptation pretrained on 1.26 million compounds.
- Combines pretrained representations with Bayesian neural networks for uncertainty estimation.
Active Learning Cycle:
- Starts with small initial labeled dataset (≈100 molecules).
- Uses Bayesian acquisition functions (BALD, EPIG) to select informative samples from unlabeled pool.
- Iteratively incorporates newly labeled data and retrains model.
- Evaluates performance using Expected Calibration Error (ECE) measurements.
This framework disentangles representation learning from uncertainty estimation, proving particularly valuable in low-data scenarios common early-stage drug discovery [9].
Ensemble Model Experimental Protocol
The ensemble approach combines BERT, RoBERTa, and XLNet without extensive pretraining requirements [25]:
- Model Integration: Implements ensemble learning with BERT, RoBERTa, and XLNet architectures.
- Training Strategy: Uses supervised fine-tuning rather than extensive pretraining from scratch.
- Resource Optimization: Specifically designed to address computational resource limitations in experimental settings.
- Performance Validation: Demonstrates significant effectiveness compared to existing advanced models while maintaining resource efficiency.
Visualizing Experimental Workflows
GEO-BERT 3D Molecular Representation Workflow
Diagram 1: GEO-BERT 3D Molecular Representation Workflow
Bayesian Active Learning with Pretrained BERT
Diagram 2: Bayesian Active Learning with Pretrained BERT
Table 2: Key Research Reagent Solutions for BERT-based Molecular Property Prediction
| Resource Category | Specific Tool/Dataset | Function and Application | Access Information |
|---|---|---|---|
| Benchmark Datasets | Tox21 Dataset [9] | Provides ≈8,000 chemical compounds with binary toxicity labels across 12 pathways; used for model validation | Publicly available |
| ClinTox Dataset [9] | Contains 1,484 FDA-approved and clinically failed drugs; evaluates clinical toxicity prediction | Publicly available | |
| Computational Frameworks | GEO-BERT Model [4] | Geometry-aware BERT for molecular property prediction; integrates 3D structural information | Open-source (GitHub: drug-designer/GEO-BERT) |
| HuggingFace Transformers [23] | Provides libraries for training and testing masked language models in Python | Open-source | |
| Pretrained Models | MolBERT [9] | BERT model pretrained on 1.26 million compounds; enables transfer learning | Reference implementation available |
| Evaluation Metrics | Expected Calibration Error (ECE) [9] | Measures reliability of uncertainty estimates in Bayesian active learning | Standard implementation |
The adaptation of BERT architectures for molecular property prediction represents a significant advancement in computational drug discovery, offering substantial improvements over traditional QSPR methods. GEO-BERT demonstrates the value of incorporating 3D structural information through its successful identification of novel DYRK1A inhibitors [4]. The integration of pretrained BERT with Bayesian active learning establishes a paradigm for data-efficient screening, reducing experimental iterations by 50% while maintaining predictive accuracy [9]. For resource-constrained environments, ensemble approaches provide a balanced solution that delivers competitive performance without extensive pretraining requirements [25]. These frameworks collectively highlight the transformative potential of adapted BERT architectures in accelerating early-stage drug discovery, enabling more efficient exploration of chemical space, and ultimately reducing the time and cost associated with identifying promising therapeutic candidates.
The application of BERT (Bidirectional Encoder Representations from Transformers) architectures has marked a significant evolution in molecular property prediction, a core task in modern drug discovery and materials science. These models, pre-trained on vast corpora of chemical data, leverage self-supervised learning to generate rich molecular representations that can be fine-tuned for specific predictive tasks with limited labeled data. The transition from traditional machine learning methods to sophisticated deep learning frameworks like BERT has been driven by the need for more accurate, efficient, and generalizable models in chemical research [9] [26]. This shift is particularly relevant in the context of materials property prediction, where the ability to accurately predict molecular behavior can dramatically reduce the time and cost associated with traditional experimental methods [27].
The fundamental advantage of BERT-based models lies in their bidirectional nature, which allows them to process molecular representations in context from both directions, capturing complex chemical patterns that unidirectional models might miss. Inspired by breakthroughs in natural language processing, chemical BERT models treat molecular structures as a "language" with its own syntax and grammar, whether represented as SMILES strings, molecular graphs, or other notation systems [26] [28]. This approach has proven particularly valuable in addressing the pervasive challenge of data scarcity in chemical research, where labeled experimental data is often limited due to the high costs and time requirements of wet lab experiments [9] [27].
Comparative Analysis of Chemical BERT Models
Model Architectures and Methodologies
Chemical BERT models share a common foundation but diverge in their architectural specifics, training methodologies, and molecular representations. The table below summarizes the key characteristics of prominent models in this domain.
Table 1: Architectural Overview of Key Chemical BERT Models
| Model Name | Core Architecture | Molecular Representation | Pre-training Strategy | Key Innovations |
|---|---|---|---|---|
| MolBERT [9] | Transformer-based BERT | SMILES strings | Masked language modeling on 1.26 million compounds | Effective disentanglement of representation learning and uncertainty estimation |
| GEO-BERT [4] | Geometry-enhanced BERT | 3D molecular conformations | Incorporates 3D positional information | Introduces atom-atom, bond-bond, and atom-bond positional relationships |
| MolLLMKD [27] | LLM-enhanced framework | 2D molecular graphs + semantic prompts | Multi-level knowledge distillation with reinforcement learning | Integrates LLM-generated prompts with graph neural networks |
| Graph Transformers [29] | Graph transformer | Molecular graphs | Masked atom prediction and property prediction | Extends self-attention to graphs with distance-aware mechanisms |
Performance Benchmarking
Rigorous evaluation across standardized benchmarks is essential for comparing model capabilities. The following table summarizes quantitative performance metrics for key chemical BERT models across various tasks.
Table 2: Performance Comparison of Chemical BERT Models on Benchmark Tasks
| Model | Tox21 AUC | ClinTox AUC | QM9 MAE | Virtual Screening Efficiency | Data Efficiency |
|---|---|---|---|---|---|
| MolBERT [9] | ~0.85 | ~0.90 | - | 50% fewer iterations for toxic compound identification | High (effective with limited labeled data) |
| GEO-BERT [4] | - | - | - | Identified two potent DYRK1A inhibitors (IC50: <1 μM) | - |
| MolLLMKD [27] | - | - | - | - | State-of-the-art on 12 benchmark datasets |
| Traditional Fingerprints (ECFP) [29] | Comparable to neural models | Comparable to neural models | - | - | - |
Recent benchmarking studies have revealed surprising insights about chemical BERT models. A comprehensive evaluation of 25 pretrained molecular embedding models across 25 datasets found that nearly all neural models showed negligible or no improvement over the traditional ECFP molecular fingerprint baseline [29]. This finding raises important questions about evaluation rigor in the field and suggests that the reported advantages of some complex models may be less pronounced than initially claimed when evaluated under standardized conditions.
Diagram 1: Chemical BERT Model Ecosystem showing the relationship between molecular representations, pre-training objectives, model variants, and downstream prediction tasks.
Experimental Protocols and Benchmarking Methodologies
Standardized Evaluation Frameworks
The assessment of chemical language models requires rigorous, standardized protocols to ensure comparable and reproducible results. Key benchmarking frameworks include:
ChemBench: An automated framework for evaluating chemical knowledge and reasoning abilities of LLMs, containing over 2,700 question-answer pairs across diverse chemistry topics. This benchmark measures reasoning, knowledge, and intuition across undergraduate and graduate chemistry curricula, with human expert performance for comparison [30].
Tox21 and ClinTox Protocols: Standardized datasets and splitting strategies for evaluating toxicology predictions. The Tox21 dataset contains approximately 8,000 compounds with binary labels across 12 toxicity pathways, while ClinTox includes 1,484 FDA-approved and failed drugs. Standard practice employs scaffold splitting with 80:20 ratio to create distinct training and testing sets, ensuring models are evaluated on structurally distinct molecules [9].
MOSES and GuacaMol: Platforms for measuring the quality, diversity, and fidelity of generated molecules, assessing the ability of models to explore chemical space effectively. These benchmarks provide standardized metrics for comparing generative model performance [26].
Data Efficiency and Active Learning Protocols
A critical advantage of BERT-based models is their performance in data-scarce environments, which is common in chemical research. Experimental protocols for evaluating data efficiency typically involve:
Bayesian Active Learning: A principled framework that quantifies the utility of conducting experiments. The Bayesian Active Learning by Disagreement (BALD) acquisition function selects samples that maximize information gain about model parameters, while Expected Predictive Information Gain (EPIG) prioritizes samples expected to most improve predictive performance [9].
Progressive Sampling: Experiments where models are trained with progressively larger subsets of available data to measure learning efficiency. MolBERT demonstrated equivalent toxic compound identification with 50% fewer iterations compared to conventional active learning, highlighting its data efficiency [9].
Diagram 2: Active Learning Workflow for Data-Efficient Molecular Property Prediction showing the iterative process of model training, uncertainty estimation, and selective sample acquisition.
Successful implementation of chemical BERT models requires familiarity with key datasets, software tools, and computational resources. The following table outlines essential components of the molecular property prediction toolkit.
Table 3: Essential Research Reagents and Computational Tools for Chemical BERT Implementation
| Resource | Type | Primary Function | Application Context |
|---|---|---|---|
| Tox21 Dataset [9] | Chemical Dataset | Benchmark for toxicity prediction | Contains ~8,000 compounds with 12 toxicity pathway assays |
| ClinTox Dataset [9] | Chemical Dataset | Distinguishes FDA-approved from failed drugs | 1,484 compounds with clinical trial toxicity outcomes |
| ZINC Database [31] | Compound Library | Source of drug-like molecules for training | Provides commercially available compounds for virtual screening |
| SMILES Notation [26] | Molecular Representation | Text-based molecular encoding | Standard input format for sequence-based models like MolBERT |
| Molecular Graphs [26] | Molecular Representation | Graph-based molecular encoding | Nodes (atoms) and edges (bonds) for graph neural networks |
| ECFP Fingerprints [29] | Molecular Representation | Circular substructure fingerprints | Traditional baseline for molecular machine learning |
| OPSIN Tool [31] | Cheminformatics Software | IUPAC name parsing | Validates chemical name-to-structure conversions |
| Scaffold Splitting [9] | Data Splitting Method | Ensures evaluation on distinct molecular scaffolds | Prevents data leakage and tests generalization capability |
Future Directions and Research Opportunities
The field of chemical BERT models continues to evolve rapidly, with several promising research directions emerging:
Multimodal Integration: Future models will likely combine molecular structure with diverse data types, including scientific literature, experimental protocols, and spectral data. The development of "active" environments where LLMs interact with tools and data, rather than merely responding to prompts, represents a significant frontier [32] [33].
3D Structural Incorporation: While models like GEO-BERT have begun incorporating 3D conformational information, more sophisticated integration of spatial and dynamic molecular properties remains an open challenge. The high computational cost of 3D conformation generation currently limits widespread application [4] [29].
Reasoning Capabilities: Recent "reasoning models" such as OpenAI's o3-mini have demonstrated substantially improved chemical reasoning capabilities, correctly answering 28%-59% of questions on the ChemIQ benchmark compared to just 7% for GPT-4o [31]. This suggests that enhanced reasoning architectures will play a crucial role in future chemical AI systems.
Evaluation Rigor: The surprising performance of traditional fingerprints against sophisticated neural models highlights the need for more rigorous evaluation standards. Future research must address this benchmarking gap to ensure meaningful progress [29].
As chemical BERT models mature, they are poised to transform materials property prediction from a largely empirical process to a more rational, accelerated workflow—ultimately reducing the time and cost associated with traditional experimental approaches while expanding the explorable chemical space for drug discovery and materials design.
Building Predictive Power: Methodologies and Real-World Applications of BERT
The application of BERT architecture to molecular property prediction represents a significant evolution in cheminformatics, transitioning from traditional descriptor-based methods to sophisticated deep-learning models. Inspired by breakthroughs in natural language processing (NLP), researchers have adapted transformer-based models to interpret chemical structures as a specialized language, where sequences like SMILES (Simplified Molecular Input Line Entry System) serve as sentences and atoms or functional groups as words [34] [35]. This approach allows models to learn rich, contextual molecular representations from massive unlabeled datasets, capturing complex structural patterns and chemical rules without costly experimental data. The core premise is that pretraining on diverse chemical corpora enables models to develop fundamental chemical intuition, which can then be efficiently fine-tuned for specific property prediction tasks with limited labeled data [9] [36]. Within the broader thesis of BERT architecture for materials property prediction, these molecular pretraining strategies demonstrate how transfer learning can address data scarcity, improve generalization, and accelerate discovery timelines in pharmaceutical research and development.
Comparative Analysis of Molecular Pretraining Approaches
Molecular pretraining strategies have diversified significantly, each employing distinct architectural choices and learning objectives to capture chemical information. The following table summarizes major approaches and their performance characteristics.
Table 1: Comparison of Molecular Pretraining Strategies and Performance
| Model | Architecture | Pretraining Strategy | Key Innovation | Reported Performance Advantages |
|---|---|---|---|---|
| Standard BERT [9] | Transformer (SMILES) | Masked Language Modeling (MLM) | Basic molecular string representation | 50% fewer iterations needed for equivalent toxic compound identification on Tox21/ClinTox vs. conventional active learning [9] |
| MLM-FG [35] | Transformer (SMILES) | Functional Group-targeted Masking | Selectively masks chemically significant functional groups | Outperformed existing SMILES & graph models in 9/11 benchmark tasks; surpassed some 3D-graph models [35] |
| GEO-BERT [4] | Transformer (3D Graph) | MLM with 3D Geometry | Incorporates atom-atom, bond-bond, and atom-bond positional relationships | Demonstrated optimal performance on multiple benchmarks; successfully identified novel DYRK1A inhibitors (IC50: <1 μM) [4] |
| MoleVers [36] | Branching Encoder | Two-Stage: Self-supervised + Auxiliary Labels | Combines masked atom prediction, dynamic denoising, and inexpensive computational labels | SOTA on 20/22 low-data MPPW benchmark datasets; ranks second on remaining two [36] |
| ECFP (Baseline) [29] | Fixed Fingerprint | Rule-based substructure identification | Traditional circular fingerprint | Extensive benchmarking (25 models, 25 datasets) showed nearly all neural models had negligible or no improvement over ECFP baseline [29] |
The experimental data reveals several key trends. First, specialized masking strategies that incorporate chemical knowledge, such as MLM-FG's functional group masking, consistently outperform standard masked language modeling [35]. Second, the integration of 3D structural information, as demonstrated by GEO-BERT, provides significant performance gains by capturing spatial relationships critical to molecular properties and interactions [4]. Third, hybrid pretraining frameworks that combine multiple objectives—such as MoleVers' integration of self-supervised and supervised pretraining—show remarkable effectiveness in data-scarce scenarios common in real-world drug discovery [36].
However, a crucial critical perspective emerges from recent benchmarking studies. A comprehensive evaluation of 25 pretrained models across 25 datasets revealed that nearly all neural approaches showed negligible or no improvement over the traditional ECFP fingerprint baseline, with only the CLAMP model (also fingerprint-based) achieving statistically significant superiority [29]. This finding raises important concerns about evaluation rigor in the field and suggests that the reported advantages of complex pretraining strategies require careful validation against simpler baselines.
Experimental Protocols and Methodologies
Data Preparation and Pretraining Corpus
Successful molecular pretraining begins with curating large-scale, diverse chemical datasets. Common sources include PubChem (containing over 100 million purchasable compounds), ZINC, and ChEMBL [35] [36]. The standard protocol involves extracting SMILES strings or 2D/3D molecular graphs from these databases. For SMILES-based models, data preprocessing includes canonicalization (standardizing string representation) and tokenization, which can occur at the character level (individual atoms, bonds) or substructure level (using a learned vocabulary or chemically aware fragmentation) [34] [35]. For graph-based approaches, molecules are represented as topological graphs with atoms as nodes and bonds as edges, often with additional features for atom type, charge, hybridization, and bond type [4] [29].
Critical to evaluating generalization is the data splitting strategy. While random splitting is common, scaffold splitting—which partitions molecules based on their core Bemis-Murcko scaffolds—provides a more rigorous test by ensuring structurally distinct molecules appear in training and test sets [9] [35]. This method prevents artificially inflated performance from evaluating on molecules structurally similar to training examples and better simulates real-world drug discovery where novel scaffolds are frequently sought.
Detailed Pretraining Methodologies
Table 2: Core Pretraining Objectives and Their Implementation
| Pretraining Objective | Mechanism | Chemical Knowledge Encoded | Implementation Example |
|---|---|---|---|
| Masked Language Modeling (MLM) | Randomly masks tokens in SMILES string; model predicts masked tokens | Contextual relationships between atoms/substructures in molecular sequences | Standard BERT: 15% masking rate; predicts original vocabulary tokens [9] |
| Functional Group Masking (MLM-FG) | Identifies and masks subsequences corresponding to functional groups | Critical chemical substructures (e.g., carboxylic acids, esters) determining molecular properties | MLM-FG: Parses SMILES, identifies functional groups via RDKit, masks 15% of FG tokens [35] |
| 3D Geometry Integration | Incorporates spatial distance/angle relationships between atoms | Three-dimensional molecular conformation critical for binding and activity | GEO-BERT: Uses atom-atom, bond-bond, atom-bond positional encodings from 3D conformers [4] |
| Dynamic Denoising | Adds noise to atom coordinates; model learns to denoise | Molecular force fields and structural stability principles | MoleVers: Applies Gaussian noise to coordinates; model predicts original equilibrium structure [36] |
| Two-Stage Pretraining | Stage 1: Self-supervised learning; Stage 2: Predicting computational labels | Transfers knowledge from inexpensive computational properties (e.g., DFT) to experimental properties | MoleVers: Stage 1: Masked atom prediction + denoising; Stage 2: Fine-tunes on auxiliary computational labels [36] |
The workflow for implementing these pretraining strategies follows a systematic pipeline, visualized below.
Diagram 1: Molecular Pretraining Workflow
Evaluation Metrics and Benchmarking
Standardized evaluation is critical for comparing pretraining approaches. For classification tasks (e.g., toxicity prediction, activity classification), the Area Under the Receiver Operating Characteristic Curve (AUC-ROC) is the primary metric, measuring the model's ability to distinguish between positive and negative classes across threshold settings [9] [35]. For regression tasks (e.g., predicting binding affinity, solubility), Mean Absolute Error (MAE) and Root Mean Squared Error (RMSE) quantify the deviation between predicted and experimental values [35].
Beyond predictive accuracy, Expected Calibration Error (ECE) measures how well the model's confidence scores align with actual accuracy, which is crucial for active learning applications where uncertainty estimation guides experimental design [9]. Benchmark datasets from MoleculeNet—including Tox21, ClinTox, HIV, BBBP, and others—provide standardized evaluation platforms [9] [35] [29]. Recent benchmarks like the Molecular Property Prediction in the Wild (MPPW) dataset, comprising 22 small datasets from ChEMBL with 50 or fewer training labels, better simulate real-world data scarcity [36].
The Scientist's Toolkit: Essential Research Reagents
Implementing molecular pretraining strategies requires both computational tools and chemical knowledge resources. The following table details essential "research reagents" for conducting these experiments.
Table 3: Essential Research Reagents for Molecular Pretraining Experiments
| Resource Category | Specific Tools / Databases | Function in Pretraining Research |
|---|---|---|
| Chemical Databases | PubChem, ZINC, ChEMBL | Provide large-scale unlabeled molecular datasets for pretraining; source of experimental labels for fine-tuning [35] [36] |
| Cheminformatics Toolkits | RDKit, OpenBabel | Process molecular representations; convert between formats; identify functional groups; generate descriptors [35] |
| Deep Learning Frameworks | PyTorch, TensorFlow, DeepGraphLibrary | Implement transformer and GNN architectures; manage pretraining and fine-tuning workflows [9] [35] |
| Molecular Representation Libraries | SMILES, SELFIES, Molecular Graphs | Standardized formats for representing chemical structures as model inputs [34] [35] |
| Benchmarking Suites | MoleculeNet, MPPW | Standardized datasets and evaluation protocols for comparing model performance [35] [36] [29] |
| Pretrained Models | GEO-BERT, MLM-FG, MoleVers | Available model weights for transfer learning; baselines for comparative studies [35] [36] [4] |
The relationship between these resources in a typical research workflow is illustrated below, showing how data flows from raw chemicals to validated predictions.
Diagram 2: Research Resource Integration
The pretraining landscape for molecular property prediction demonstrates a clear evolution from generic masked language modeling toward chemically-aware strategies that explicitly incorporate structural knowledge. Approaches that target functionally important substructures (MLM-FG), integrate 3D geometry (GEO-BERT), or combine multiple pretraining objectives (MoleVers) show consistent performance advantages across standardized benchmarks [35] [36] [4]. The integration of these pretrained models with active learning frameworks further enhances their practical utility, enabling more efficient experimental design and compound prioritization in drug discovery pipelines [9].
However, the field faces critical challenges regarding evaluation rigor and practical utility. The surprising benchmarking result that most neural approaches fail to consistently outperform traditional fingerprints raises important questions about the true extent of progress in this domain [29]. Future research should prioritize (1) more rigorous evaluation against simple baselines, (2) standardization of benchmarking protocols to prevent data leakage, and (3) development of pretraining strategies that more effectively capture the fundamental principles of molecular structure-activity relationships. For researchers and drug development professionals, the current evidence suggests adopting a hybrid approach that leverages the strengths of both modern pretrained models and traditional chemical descriptors, while maintaining realistic expectations about the achievable performance gains in practical applications.
The accurate prediction of materials and molecular properties is a cornerstone of modern drug development and materials science. However, the field consistently grapples with the fundamental challenge of data sparsity; high-quality, annotated experimental data is often scarce and costly to obtain, creating a significant bottleneck for training robust machine learning models [37]. Within the broader context of BERT architecture research for materials property prediction, two innovative strategies have emerged as powerful solutions: multitask learning (MTL) and SMILES enumeration. Multitask learning improves generalization by leveraging information from multiple related tasks, thereby effectively amplifying the learning signal from limited data [38] [39]. Concurrently, SMILES enumeration acts as a powerful data augmentation technique, expanding the effective size of training sets by representing a single molecule with multiple valid text strings [40]. This guide provides an objective comparison of these approaches, detailing their experimental protocols, performance, and practical utility for researchers and scientists.
Multitask learning is a subfield of machine learning where multiple learning tasks are solved simultaneously, exploiting commonalities and differences across tasks to improve generalization and prediction accuracy for each individual task [39]. The central idea is that by learning tasks in parallel using a shared representation, the model can prevent overfitting and perform better on sparse data tasks. As Rich Caruana stated in his seminal 1997 work, MTL "improves generalization by using the domain information contained in the training signals of related tasks as an inductive bias" [39].
Key Methodologies and Optimization Approaches
Several methodological frameworks have been developed to implement MTL effectively:
- Task Grouping and Overlap: Information can be shared selectively across tasks. Tasks may be grouped in a hierarchy or related according to a learned metric, where similarity in a underlying parameter basis indicates relatedness [39].
- Multi-task Optimization: This is inherently a multi-objective optimization problem. Modern approaches include Multi-task Bayesian Optimization, which builds multi-task Gaussian process models to capture inter-task dependencies, and Evolutionary Multi-tasking, which uses population-based search algorithms to progress multiple optimization tasks simultaneously through genetic transfer [39].
- Direct Metric Optimization: Some methods, such as those based on the Alternating Direction Method of Multipliers (ADMM), directly optimize evaluation metrics for a family of MTL problems by combining a regularizer on the weight matrix with a sum of structured hinge losses [41].
Experimental Evidence in Materials Science
The PolyQT (Polymer Quantum-Transformer) model exemplifies a sophisticated MTL approach applied to polymer informatics. This hybrid architecture combines Quantum Neural Networks (QNNs) with a Transformer to address sparse data challenges [37]. In prediction experiments for six key polymer properties, the PolyQT model demonstrated significant advantages, achieving R² values of 0.85, 0.77, 0.85, 0.83, and 0.92 for ionization energy, dielectric constant, glass transition temperature, refractive index, and polymer density, respectively, outperforming all benchmarked classical models [37]. Crucially, its performance remained robust under different data sparsity conditions (40%, 60%, and 80% data), confirming MTL's utility in data-limited scenarios [37].
SMILES Enumeration: Augmenting Molecular Representations
SMILES (Simplified Molecular-Input Line-Entry System) is a line notation for representing molecular structures as text strings. A single molecule can be represented by multiple, equally valid SMILES strings due to different possible atom ordering during the traversal of the molecular graph [40]. SMILES enumeration, also known as randomized SMILES, leverages this property as a powerful data augmentation technique.
Implementation and Impact on Model Generalization
In practice, models are trained using different SMILES representations of the same molecule for each epoch. For example, a model trained on one million molecules for 300 epochs would be exposed to approximately 300 million different randomized SMILES, vastly increasing the effective diversity of the training data [40]. Benchmark studies have conclusively shown that models trained on randomized SMILES generalize better than those trained on canonical (unique) SMILES. They generate chemical spaces that are more uniform, complete, and closed, representing the target chemical space more accurately [40].
A particularly counter-intuitive yet profound finding is that the ability of language models to generate invalid SMILES is actually beneficial rather than detrimental [42]. Research demonstrates that invalid SMILES are typically sampled with significantly lower likelihoods than valid SMILES, meaning that filtering them out acts as an intrinsic self-corrective mechanism that removes low-quality samples [42]. Enforcing 100% validity, as done with alternative representations like SELFIES, can introduce structural biases and impair a model's ability to learn the true data distribution and generalize to unseen chemical space [42].
Performance Comparison: Quantitative Benchmarks
The following tables summarize experimental data comparing the performance of these and other related approaches on standardized benchmarks.
Table 1: Performance Comparison of Cross-Modal Knowledge Transfer on LLM4Mat-Bench (Selected Tasks)
| Predictive Task | SOTA Existing Model (MAE) | SOTA Presented Model (MAE) | Performance Boost | Best-Performing Architecture |
|---|---|---|---|---|
| Formation Energy (FEPA) | MatBERT-109M: 0.126 | 0.11488 ± 0.00018 | +8.8% | imKT@ModernBERT [5] |
| Band Gap (OPT) | MatBERT-109M: 0.235 | 0.1985 ± 0.0019 | +15.5% | imKT@BERT [5] |
| Total Energy | MatBERT-109M: 0.194 | 0.1172 ± 0.0005 | +39.6% | imKT@ModernBERT [5] |
| Band Gap (MBJ) | MatBERT-109M: 0.491 | 0.3773 ± 0.0030 | +23.2% | imKT@ModernBERT [5] |
| Exfoliation Energy | MatBERT-109M: 37.445 | 29.5 ± 1.4 | +21.2% | imKT@RoFormer [5] |
Table 2: Performance of PolyQT (MTL) vs. Benchmark Models on Polymer Properties
| Property Predicted | PolyQT (R²) | Best Benchmark Model (R²) | Key Advantage |
|---|---|---|---|
| Ionization Energy | 0.85 | <0.85 (TransPolymer, NN, etc.) | Superior accuracy [37] |
| Dielectric Constant | 0.77 | <0.77 | Superior accuracy [37] |
| Glass Transition Temp. | 0.85 | <0.85 | Superior accuracy [37] |
| Refractive Index | 0.83 | <0.83 | Superior accuracy [37] |
| Polymer Density | 0.92 | <0.92 | Superior accuracy [37] |
Table 3: Impact of SMILES Enumeration on Model Generalization
| Training Method | % of GDB-13 Generated | Validity Rate | Distribution Matching | Key Finding |
|---|---|---|---|---|
| Canonical SMILES | ≤68% | ~99.9% | Lower | Suboptimal coverage [40] |
| Randomized SMILES | Up to ~100% | ~90.2% | Higher (Better Fréchet ChemNet Distance) | Better representation of target space [40] [42] |
Experimental Protocols and Workflows
Protocol for Cross-Modal Knowledge Transfer (for Materials Property Prediction)
This protocol, derived from state-of-the-art research, involves transferring knowledge from structure-aware models to composition-based models [5].
- Pretraining a Multimodal Foundation Model: Begin by pretraining a model (e.g., MultiMat) contrastively on multiple modalities of materials data, such as crystal structure, density of electronic states, charge density, and textual description [5].
- Chemical Language Model (CLM) Alignment (Implicit Knowledge Transfer):
- Train a CLM (e.g., a BERT variant) on a large corpus of chemical compositions via Masked Language Modeling (MLM).
- Align the embedding space of the CLM with the embeddings from the pretrained multimodal foundation model. This transfers structural and electronic knowledge to the composition-based CLM without explicitly generating structures [5].
- Fine-tuning and Evaluation:
Protocol for Training with SMILES Enumeration (for Molecular Property Prediction)
This protocol outlines the use of randomized SMILES for data augmentation [40].
- Data Preparation and Tokenization:
- Obtain a dataset of molecules (e.g., from ChEMBL or GDB-13) represented as canonical SMILES.
- Implement an atom-order randomization routine to generate multiple non-unique SMILES strings for each molecule. An "unrestricted" version that avoids built-in traversal fixes can yield greater diversity [40].
- Tokenize the SMILES strings on a character basis, with special handling for multi-character tokens like "Cl", "Br", and ring indices above 9 [40].
- Model Training with Epoch-Wise Enumeration:
- For each training epoch, generate a new set of randomized SMILES for every molecule in the training set. This ensures the model sees a vast number of string variations without increasing the number of unique molecules [40].
- Use a standard architecture like an LSTM-based RNN or Transformer. The input is fed through an embedding layer, followed by recurrent/attention layers and a final linear layer with softmax to predict the next token [40] [42].
- Employ the "teacher's forcing" strategy and minimize the average negative log-likelihood (NLL) of the tokenized SMILES strings across the batch [40].
- Sampling and Post-Processing:
- Sample new SMILES strings from the trained model.
- Filter out invalid SMILES. Crucially, this step also filters low-likelihood samples, improving the overall quality of the generated set [42].
Workflow and Conceptual Diagrams
The Scientist's Toolkit: Essential Research Reagents
Table 4: Key Computational Tools and Datasets for Overcoming Data Limits
| Tool / Resource | Type | Primary Function | Relevance to Data Sparsity |
|---|---|---|---|
| Matbench [43] | Benchmark Suite | Standardized set of 13 ML tasks for inorganic materials. | Provides reliable, pre-cleaned datasets for fair model comparison and evaluation of generalization. |
| LLM4Mat-Bench [44] | Benchmark Suite | Largest benchmark for evaluating LLMs on crystalline materials properties (~1.9M structures). | Enables scalable testing of models across 45 distinct properties and different input modalities. |
| Automatminer [43] | Automated ML Pipeline | End-to-end pipeline for materials property prediction from primitives. | Serves as a powerful baseline and reference algorithm, automating feature generation and model selection. |
| Randomized SMILES | Data Augmentation | Algorithm for generating multiple SMILES representations per molecule. | Directly increases effective training data size, improving model robustness and generalization [40]. |
| Multi-task Gaussian Process [39] | Optimization Model | Bayesian model for capturing inter-task dependencies. | Facilitates knowledge transfer between related tasks in an MTL setting, improving data efficiency. |
| Quantum-Transformer (PolyQT) [37] | Model Architecture | Hybrid model combining Quantum Neural Networks and Transformer. | Designed to capture complex, nonlinear relationships in sparse polymer datasets. |
In the field of materials informatics, a significant challenge persists: how to accurately predict the properties of a material when only its chemical composition is known, and its precise crystal structure remains undetermined. Structure-aware models, such as crystal graph neural networks (GNNs), have demonstrated excellent performance on experimentally synthesized compounds where crystallographic data is available [5]. However, their application is limited when exploring previously inaccessible domains of chemical space, a task for which structure-agnostic predictive algorithms are essential [5].
The advent of BERT-based architectures and other transformer models has revolutionized many fields, including materials science. These chemical language models (CLMs) reframe composition-based property prediction as a sequence modeling task [5]. Yet, a fundamental gap remains between the wealth of information embedded in known crystal structures and the simplicity of compositional data. Cross-modal knowledge transfer has emerged as a powerful strategy to bridge this divide, enabling the transfer of knowledge from data-rich modalities (like crystal structures) to improve predictions in data-scarce modalities (like chemical compositions alone).
This guide provides a comparative analysis of the leading cross-modal knowledge transfer approaches for materials property prediction, detailing their experimental protocols, performance benchmarks, and implementation requirements to assist researchers in selecting appropriate methodologies for their specific applications.
Comparative Analysis of Cross-Modal Transfer Approaches
Performance Benchmarking
The following table summarizes the experimental performance of major cross-modal knowledge transfer approaches compared to established baseline methods across key materials property prediction tasks.
Table 1: Performance Comparison of Cross-Modal Knowledge Transfer Approaches
| Method | Architecture Type | Key Properties Predicted | Performance Metrics | Dataset(s) | Compared Baselines |
|---|---|---|---|---|---|
| Implicit Knowledge Transfer (imKT) [5] | Chemical Language Model (ModernBERT, RoFormer) | Formation energy per atom (FEPA), Band gap (OPT), Total energy | MAE: 0.11488 (FEPA, +8.8% improvement), 0.1985 (Band gap, +15.5%), 0.1172 (Total energy, +39.6%) | LLM4Mat-Bench, MatBench | MatBERT-109M, Gemma2-9b-it, LLM-Prop-35M |
| Explicit Knowledge Transfer (exKT) [5] | LLM Crystal Structure Predictor + GNN | Properties requiring structural knowledge | State-of-the-art in 25/32 benchmark tasks [5] | LLM4Mat-Bench | Structure-agnostic baselines |
| CroMEL [45] | Cross-modality material embedding loss | Experimentally measured formation enthalpies, Band gaps | R² > 0.95 for formation enthalpies and band gaps [45] | 14 experimental materials datasets | Conventional machine learning |
| PolyQT [37] | Quantum-Transformer Hybrid | Ionization energy, Dielectric constant, Glass transition temperature | R²: 0.85 (Ionization Energy), 0.77 (Dielectric Constant), 0.85 (Glass Transition Temp.) | 6 polymer datasets | Gaussian Processes, Neural Networks, Random Forests |
Methodology Comparison
Table 2: Technical Comparison of Cross-Modal Transfer Methodologies
| Method | Transfer Mechanism | Modalities Bridged | Training Complexity | Data Requirements | Key Advantages |
|---|---|---|---|---|---|
| Implicit Transfer (imKT) [5] | Embedding alignment via contrastive pretraining | Composition → Multimodal embeddings (structure, DOS, charge density, text) | High (multimodal pretraining) | Large source dataset for pretraining | Direct property prediction, no explicit structure generation |
| Explicit Transfer (exKT) [5] | Crystal structure generation followed by property prediction | Composition → Crystal structure → Property | Very High (two-stage training) | Structure-property pairs for training | Leverages powerful structure-aware GNNs |
| CroMEL [45] | Distribution alignment via Wasserstein distance | Calculated crystal structures → Experimental compositions | Medium (embedding alignment) | Paired composition-structure data | Handles polymorphic crystal structures effectively |
| PolyQT [37] | Quantum-classical feature fusion | SMILES representations → Quantum-enhanced embeddings | Very High (quantum-classical hybrid) | Polymer SMILES and property data | Superior performance on sparse data |
Experimental Protocols and Workflows
Implicit Cross-Modal Knowledge Transfer (imKT)
The implicit knowledge transfer approach eliminates the need for explicit structure generation by aligning compositional representations with multimodal embeddings [5].
Workflow Description: The process begins with chemical language models (CLMs) initially pretrained using masked language modeling (MLM) on extensive materials science text corpora. The core transfer mechanism involves aligning these CLM embeddings with those from a foundation model (MultiMat) that was contrastively pretrained on four distinct materials modalities: crystal structure, density of electronic states, charge density, and textual description [5]. This alignment creates a shared embedding space where compositional information is infused with structural knowledge without explicit structure prediction. The aligned model can then be fine-tuned on specific property prediction tasks using standard regression or classification heads.
Explicit Cross-Modal Knowledge Transfer (exKT)
Explicit knowledge transfer employs a two-stage process where crystal structures are first generated from compositions before property prediction [5].
Workflow Description: This methodology uses large language models, such as CrystaLLM, specifically trained for crystal structure prediction from chemical compositions [5]. In the first stage, the LLM generates probable crystal structures given input stoichiometries. These generated structures then serve as input to structure-aware predictors, typically graph neural networks (GNNs) that have been pretrained on established structure-property datasets. The GNNs process the crystal graphs, incorporating information about atomic arrangements, bond lengths, and coordination environments to predict target properties. This approach effectively transfers knowledge from the structural domain to enhance composition-based prediction.
Cross-Modality Material Embedding Loss (CroMEL)
CroMEL implements a probabilistic approach to align embedding distributions across different material modalities [45].
Workflow Description: CroMEL addresses the challenge of transferring knowledge from calculated crystal structures to experimental compositions where structural data is unavailable. The method employs two encoders: a structure encoder (π) trained on source datasets with crystal structures, and a composition encoder (ψ) that processes chemical compositions [45]. The core innovation is the cross-modality material embedding loss, which minimizes the statistical divergence (using Wasserstein distance) between the probability distributions of structure embeddings and composition embeddings. This alignment ensures that the composition encoder captures latent structural information, enabling effective knowledge transfer even without explicit structure prediction.
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Computational Tools and Resources for Cross-Modal Materials Research
| Tool/Resource | Type | Function/Role | Access/Implementation |
|---|---|---|---|
| MultiMat Foundation Model [5] | Multimodal Embedding Model | Provides aligned representations across crystal structure, DOS, charge density, and text | Research implementation required |
| CrystaLLM [5] | Large Language Model | Generates crystal structures from chemical compositions | Research implementation required |
| CroMEL Framework [45] | Loss Function/Algorithm | Aligns embedding distributions across material modalities | Custom implementation based on published criteria |
| PolyQT Framework [37] | Quantum-Transformer Hybrid | Enhances prediction on sparse polymer datasets | Requires quantum computing resources |
| JARVIS-DFT Dataset [5] | Materials Database | Benchmark dataset for property prediction tasks | Publicly available |
| MatBench [5] | Benchmarking Suite | Standardized evaluation framework for materials informatics | Publicly available |
| LLM4Mat-Bench [5] | Benchmarking Suite | Evaluation framework for language models in materials science | Publicly available |
Cross-modal knowledge transfer represents a paradigm shift in materials property prediction, effectively bridging the critical gap between compositional and structural representations. The experimental data demonstrates that both implicit and explicit transfer approaches can significantly outperform conventional unimodal methods, achieving state-of-the-art results on standardized benchmarks.
For researchers implementing these methodologies, the choice between implicit and explicit transfer depends on specific application requirements: implicit transfer offers greater efficiency for direct property prediction, while explicit transfer provides interpretable structural intermediates. Emerging approaches like CroMEL and quantum-enhanced models show particular promise for challenging scenarios involving experimental data sparsity and complex polymer systems.
As BERT-based architectures continue to evolve in materials science, cross-modal integration will likely play an increasingly central role in enabling accurate, data-efficient exploration of chemical space and accelerating the discovery of novel materials with tailored properties.
The accurate prediction of chemical toxicity is a critical challenge in drug discovery, environmental safety, and regulatory science. Unexpected toxicities, particularly drug-induced liver injury (DILI), remain a leading cause of late-stage clinical trial failures and market withdrawals, costing the pharmaceutical industry an estimated $350 million annually per company [46]. Traditional methods, including quantitative structure-activity relationship (QSAR) models and in vitro assays, have been widely used but often struggle with generalizability, specificity, and providing mechanistic insights [46] [47].
The integration of advanced artificial intelligence (AI) techniques is creating a paradigm shift in computational toxicology. This case study objectively compares two powerful, yet philosophically distinct, AI frameworks for toxicity prediction: VitroBERT, a BERT-based model for molecular representation learning, and BATCHIE, a Bayesian active learning platform for efficient experimental design [48] [49] [50]. The analysis is framed within a broader thesis on leveraging BERT architectures for materials property prediction, demonstrating how these models address the core challenges of data scarcity, biological context integration, and translational accuracy between experimental domains.
Methodologies & Experimental Protocols
VitroBERT: Biologically Informed Molecular Representations
VitroBERT is a Bidirectional Encoder Representations from Transformers (BERT) model specifically designed to generate molecular embeddings enriched with biological context [48]. Its pretraining strategy fundamentally extends traditional unsupervised molecular representation learning.
- Model Architecture and Pretraining: The model is built on a shared BERT encoder coupled with multiple task-specific heads [48]. During pretraining, it simultaneously learns from three distinct tasks:
- A masking head that recovers masked tokens in SMILES strings, learning the underlying grammar of chemical structures.
- A physicochemical property head that predicts intrinsic molecular characteristics.
- An in vitro assay head that models biological interactions, using data from large-scale bioactivity profiles.
- Pretraining Datasets: The model was pretrained on two large-scale in vitro datasets:
- A DILI-centric in-house dataset compiled from the OFF-X database and internal ADME assays, comprising ~1.26 million compounds across ~1200 classification tasks related to liver toxicity and pharmacokinetics [48].
- The public ChEMBL20 dataset, a manually curated collection of ~445,000 drug-like molecules annotated across 1243 binary classification tasks [48].
- Fine-tuning and Loss Functions: For downstream toxicity prediction, the pretrained VitroBERT model generates embeddings for molecules, which are then used to train a lightweight multilayer perceptron (MLP) head. To address severe class imbalance common in toxicity datasets, the study rigorously evaluated loss functions, identifying weighted Focal loss as the most effective [48].
BATCHIE: Bayesian Active Learning for Combination Screening
BATCHIE (Bayesian Active Treatment Combination Hunting via Iterative Experimentation) adopts an orthogonal approach, focusing not on molecular representation but on optimizing the experimental design process itself to make combination drug screens tractable [49] [50].
- Core Algorithm: BATCHIE uses a Bayesian active learning strategy to conduct experiments dynamically in small batches [49]. Each subsequent batch is designed to be maximally informative based on the results of previous experiments, a method grounded in information theory.
- Experimental Design Criterion: The platform uses a Probabilistic Diameter-based Active Learning (PDBAL) criterion. PDBAL selects experiments that minimize the expected distance between any two posterior samples after observing the new data, ensuring theoretical near-optimality [49].
- Predictive Model: BATCHIE is compatible with any Bayesian model. The implemented model uses hierarchical Bayesian tensor factorization, which decomposes a combination's effect on a cell line into individual drug effects and interaction terms using learned embeddings for cell lines and drug-doses [49].
- Prospective Validation Protocol: The model's efficacy was validated in a real-world screen of a 206-drug library across 16 pediatric cancer cell lines. The adaptive design explored only 4% of the 1.4 million possible combinations to accurately predict synergistic drug pairs [49].
The workflow for BATCHIE is distinct from the static training of VitroBERT, as illustrated below.
Performance Comparison & Experimental Data
The two frameworks were evaluated on different, highly relevant tasks. The quantitative results from their respective studies are summarized in the table below.
Table 1: Comparative Performance of VitroBERT and BATCHIE
| Model | Primary Task | Key Metric | Reported Performance | Benchmark / Baseline | Key Advantage |
|---|---|---|---|---|---|
| VitroBERT [48] | Predicting in vivo DILI endpoints from molecular structure | Improvement in AUC (Area Under the Curve) | Up to 29% improvement in biochemistry-related tasks and 16% gain in histopathology endpoints vs. unsupervised pretraining (MolBERT). No significant gain in clinical tasks. | MolBERT (unsupervised BERT) | Embeds biological context from in vitro data into molecular representations. |
| BATCHIE [49] [50] | Large-scale combination drug screening | Experimental Efficiency & Predictive Accuracy | Accurately predicted synergistic combinations after testing only 4% of 1.4M possible experiments. Identified a panel of effective combinations for Ewing sarcoma. | Traditional fixed-design screens | Drastically reduces the experimental burden and cost of combination screens. |
Contextualizing VitroBERT's Performance
The performance of transformer-based models like VitroBERT can be further contextualized by a broader comparison against traditional molecular descriptors. A separate, comparative study on toxicity prediction provides this insight, with key data shown in the table below.
Table 2: Performance Comparison of Molecular Descriptors vs. AI Language Models on Standard Toxicity Datasets (ROC-AUC) [51]
| Model Type | Representation | Tox21 (Avg.) | ClinTox | DILIst |
|---|---|---|---|---|
| Descriptor-Based | Mordred | 0.855 | - | - |
| Descriptor-Based | RDKit | - | 0.721 | 0.620 |
| Language Model | MolBERT (SMILES) | 0.801 | - | - |
| Language Model | GPT-3 (Descriptions) | - | 0.996 | - |
| Language Model | GPT-3 (Chemical Names) | - | - | 0.806 |
This data underscores a critical insight for BERT-based property prediction research: while molecular descriptors can be robust for multi-endpoint predictions (e.g., Tox21), language models can achieve superior performance on more focused classification tasks, especially when leveraging textual chemical representations [51].
The Scientist's Toolkit: Essential Research Reagents & Materials
The experimental workflows for these AI models rely on specific data resources and computational tools. The following table details key components of the modern computational toxicologist's toolkit.
Table 3: Key Research Reagents and Resources for AI-Driven Toxicity Prediction
| Resource Name | Type | Primary Function in Research | Relevance to Model |
|---|---|---|---|
| OFF-X Database [48] | Bioactivity Database | Provides data on drug off-target effects and associations with adverse drug reactions (ADRs). | VitroBERT: Source of DILI-centric in vitro assay data for pretraining. |
| ChEMBL [48] | Bioactivity Database | A large, open-source database of bioactive molecules with drug-like properties and assay data. | VitroBERT: Public source of diverse bioactivity data for pretraining. |
| DILIrank [46] | Curated Dataset | A benchmark dataset used for training and validating DILI prediction models. | VitroBERT / ToxPredictor: Provides standardized clinical DILI labels for model evaluation. |
| Open TG-GATEs [48] [52] | Toxicogenomics Database | A comprehensive resource containing in vivo and in vitro transcriptomic and pathological data from compound treatments. | Used for training and validating various models, including histopathology endpoints for VitroBERT and as a data source for AIVIVE [52]. |
| TOXRIC [53] | Toxicology Data Platform | A comprehensive database of toxicological data and benchmarks, providing ML-ready datasets for 1,474 endpoints. | General Use: A valuable resource for obtaining standardized datasets for model training and benchmarking. |
| BATCHIE Software [49] | Computational Platform | An open-source Python package for implementing Bayesian active learning in combination drug screens. | BATCHIE: The core software implementation of the active learning framework. |
This case study demonstrates that VitroBERT and BATCHIE offer powerful, complementary solutions for different facets of the toxicity prediction problem. VitroBERT excels at learning biologically meaningful molecular representations from existing in vitro data, directly enhancing the accuracy of predicting specific in vivo toxicological endpoints like DILI [48]. Its strength lies in transferring knowledge from large-scale bioassay data to inform downstream predictive tasks, a core tenet of effective BERT-based property prediction.
In contrast, BATCHIE addresses the foundational challenge of experimental scalability. Its Bayesian active learning framework provides a statistically rigorous and highly efficient method for navigating vast experimental spaces, such as combination drug screens, with minimal resource expenditure [49] [50].
The future of AI in toxicology points toward the integration of such specialized frameworks. A promising direction is the development of multi-modal models that combine molecular representations (like those from VitroBERT) with transcriptomic data from resources like DILImap [46] or generative AI for in vitro to in vivo extrapolation (IVIVE) as seen with AIVIVE [52]. Furthermore, incorporating active learning principles from BATCHIE into the data acquisition and model training phases for molecular models could optimize the use of costly experimental resources, creating a more iterative and efficient AI-driven discovery pipeline. This synthesis of deep representation learning and optimal experimental design will be pivotal in developing more predictive, reliable, and actionable models for chemical safety assessment.
The electronic band gap is a fundamental property of crystalline materials that determines their electrical conductivity and optical characteristics, making it a critical parameter for designing semiconductors, solar cells, and other electronic devices [54] [55]. Accurate prediction of this property has long challenged materials scientists due to the complex relationship between chemical composition, crystal structure, and electronic behavior. Traditional approaches using density functional theory (DFT) calculations often suffer from the "band gap problem"—a significant discrepancy between calculated and experimental values—while also being computationally intensive and limited to materials with known crystal structures [54] [55]. This case study examines how modern computational approaches, including machine learning (ML) models and natural language processing (NLP) techniques, are transforming band gap prediction by enabling faster, more accurate estimates across diverse material classes.
Within the broader context of BERT architecture materials property prediction research, band gap prediction represents a compelling application domain where transformer-based models demonstrate significant potential. Foundation models are catalyzing a transformative shift in materials science by enabling scalable, general-purpose AI systems for scientific discovery [56]. Unlike traditional machine learning models which are typically narrow in scope, foundation models offer cross-domain generalization and exhibit emergent capabilities well-suited to materials science challenges [56]. This case study will objectively compare the performance of various computational approaches for band gap prediction, with particular attention to how BERT-inspired architectures are addressing longstanding limitations in the field.
Comparative Analysis of Band Gap Prediction Methodologies
Performance Metrics Across Prediction Approaches
Table 1: Comparison of Band Gap Prediction Methods and Their Performance
| Method Category | Specific Approach | Data Input | Key Performance Metrics | Materials Tested | Primary Advantages |
|---|---|---|---|---|---|
| Traditional DFT | PBE Functional | Crystal Structure | Systematic underestimation (~50%) | Various | Strong theoretical foundation |
| Advanced DFT | GW Approximation | Crystal Structure | High accuracy | Small systems | High accuracy for known structures |
| Machine Learning | Gradient Boosting Decision Trees (GBDT) | Composition & Elemental Features | R²: >0.950, RMSE: <0.4 eV [54] | Binary semiconductors | High accuracy with computational efficiency |
| Machine Learning | Support Vector Regression (SVR) | Composition & Elemental Features | R²: >0.950, RMSE: <0.4 eV [54] | Binary semiconductors | Strong performance with limited data |
| Machine Learning | Random Forests (RF) | Composition & Elemental Features | R²: >0.950, RMSE: <0.4 eV [54] | Binary semiconductors | Robust to feature scaling |
| Transfer Learning | Pre-trained NN + Fine-tuning | PBE gaps + Limited GW data | MAE: 0.27 eV, R: 0.97 [57] | 2D Monolayers | Addresses data scarcity for accurate methods |
| Interpretable ML | SISSO-assisted ML | Elemental Features + PBE gaps | High interpretability [54] | Binary Compounds | Physical insights alongside predictions |
| Simple Learned Model | Element-weighted ReLU | Chemical Composition Only | Not specified | Crystalline Materials | Composition-only, highly interpretable [55] |
| LLM-Based Pipeline | LLM-Prompt Extraction → ML | Literature Text → Structured Data | 19% MAE reduction vs. human-curated database [58] | Various from literature | Leverages experimental data from literature |
Methodological Approaches and Experimental Protocols
Traditional Density Functional Theory
DFT calculations represent the traditional computational approach for band gap prediction, with the Perdew-Burke-Ernzerhof (PBE) functional being widely used for high-throughput screening [57]. The fundamental protocol involves: (1) obtaining or optimizing the crystal structure; (2) performing self-consistent field calculations to determine the ground-state electron density; (3) computing the electronic band structure; and (4) extracting the band gap as the energy difference between the valence band maximum and conduction band minimum. While computationally feasible for high-throughput screening, standard DFT functionals like PBE systematically underestimate band gaps by approximately 50% due to the exchange-correlation problem [57]. More accurate methods like the GW approximation provide better agreement with experiment but require substantially more computational resources, limiting their application to smaller systems [57].
Feature-Assisted Machine Learning
Feature-assisted ML approaches combine traditional algorithms with interpretability-focused techniques like the sure independence screening and sparsifying operator (SISSO) [54]. The experimental protocol typically involves: (1) curating a dataset of known materials and their band gaps (e.g., 1,107 binary semiconductors from the Materials Project); (2) computing or gathering 23 input features including electronegativity, ionization energy, atomic radii, and PBE-calculated band gaps; (3) training multiple ML models (SVR, RF, GBDT) with three-fold cross-validation; (4) assessing feature importance using permutation importance methods; and (5) integrating top features into SISSO to derive interpretable descriptors [54]. This approach highlights the critical role of electronegativity in determining band gaps while maintaining high predictive accuracy (R² > 0.950, RMSE < 0.4 eV) [54].
Transfer Learning for Data-Efficient Prediction
Transfer learning addresses the challenge of limited high-quality band gap data by leveraging knowledge from large datasets of less accurate calculations [57]. The protocol for 2D materials involves: (1) pre-training a neural network on 2,915 non-metallic monolayers from the Computational 2D Materials Database (C2DB) with PBE-calculated band gaps; (2) using 290 compositional descriptors generated by the XENONPY package; (3) transferring the learned representations to a small dataset of GW-calculated band gaps; and (4) fine-tuning the model for optimal performance [57]. This approach achieves exceptional correlation (Pearson coefficient of 97%) and reduced MAE (0.27 eV) compared to direct machine learning, demonstrating the power of transfer learning for overcoming data scarcity [57].
Simple Composition-Based Models
Simple models using only chemical composition offer an alternative for cases where structural information is unavailable [55]. The methodology involves: (1) analyzing the empirical distribution of band gaps to frame prediction as modeling a mixed random variable; (2) designing a model with one parameter per element; (3) computing a weighted average of element parameters based on chemical formula stoichiometry; and (4) applying a ReLU activation (max(weighted average, 0)) to produce non-negative band gap predictions [55]. This approach provides heuristic chemical interpretability, with elements having greater parameters associated with larger band gaps, while requiring only compositional information [55].
LLM-Based Data Extraction and Prediction
Large language models enable the creation of specialized datasets from scientific literature for subsequent ML training [58]. The pipeline involves: (1) using LLM prompts to extract band gap data from materials science literature with an order of magnitude lower error rate than automated extraction; (2) applying additional prompts to select experimentally measured properties from pure, single-crystalline bulk materials; (3) constructing a dataset larger and more diverse than human-curated databases; and (4) training machine learning models on the extracted data [58]. This approach demonstrates a 19% reduction in mean absolute error compared to models trained on human-curated databases, highlighting the potential of LLMs to overcome data scarcity in materials science [58].
BERT Architectures in Materials Property Prediction
Adaptation of BERT for Materials Science
BERT architectures have been specifically adapted for materials science applications through models like MaterialsBERT, which was trained on 2.4 million materials science abstracts to outperform baseline models in named entity recognition tasks [59]. The adaptation process involves: (1) continued pre-training of existing BERT models (e.g., PubMedBERT) on domain-specific corpora; (2) developing custom ontologies for materials science concepts (POLYMER, PROPERTY_VALUE, etc.); (3) fine-tuning for specific tasks like property extraction; and (4) integrating extracted data into predictive modeling pipelines [59]. This approach has enabled the extraction of approximately 300,000 material property records from 130,000 abstracts, demonstrating the scalability of BERT-based information extraction for materials science [59].
Molecular Property Prediction with BERT
For molecular property prediction, BERT architectures are modified to handle chemical representations like SMILES strings [60]. The experimental protocol includes: (1) exploring various positional embeddings (absolute, relative_key, rotary) to capture structural information in molecular sequences; (2) pre-training on large datasets of unlabeled SMILES representations (∼7.9 million instances); (3) fine-tuning on downstream property prediction tasks; and (4) evaluating zero-shot learning capabilities for predicting properties of unseen molecular structures [60]. These approaches demonstrate how transformer architectures can capture complex relationships in chemical data, though their application to band gap prediction specifically remains an emerging area compared to traditional machine learning methods.
Experimental Workflow for Band Gap Prediction
The following diagram illustrates the generalized experimental workflow for machine learning-based band gap prediction, integrating elements from multiple methodologies discussed in this case study:
Diagram 1: Band Gap Prediction Workflow showing the integration of data sources and processing methods.
Research Reagent Solutions: Computational Tools for Band Gap Prediction
Table 2: Essential Computational Tools and Datasets for Band Gap Prediction Research
| Tool/Dataset Name | Type | Primary Function | Relevance to Band Gap Prediction |
|---|---|---|---|
| Materials Project | Database | Repository of computed materials properties | Source of DFT-calculated band gaps for training [54] |
| C2DB | Database | Computational 2D Materials Database | Source of PBE and GW band gaps for 2D materials [57] |
| XENONPY | Software Package | Material descriptor generator | Creates 290 compositional descriptors for ML models [57] |
| SISSO | Algorithm | Sure Independence Screening and Sparsifying Operator | Derives interpretable descriptors from feature space [54] |
| MaterialsBERT | Language Model | Domain-specific BERT for materials science | Extracts property data from scientific literature [59] |
| scikit-learn | Software Library | Machine Learning in Python | Implements SVR, RF, GBDT algorithms for prediction [54] |
| PolymerScholar | Web Interface | Exploration of extracted polymer data | Locates material property information from abstracts [59] |
| Open MatSci ML Toolkit | Infrastructure | Standardizes materials learning workflows | Supports development of foundation models [56] |
This comparative analysis demonstrates that while traditional DFT calculations provide the theoretical foundation for band gap prediction, machine learning approaches offer superior computational efficiency and, in many cases, improved accuracy, particularly when leveraging large datasets or transfer learning strategies. The emerging paradigm of using LLMs and BERT-based architectures for data extraction and property prediction shows significant promise for addressing the data scarcity challenges that have long limited materials informatics.
Within the broader context of BERT architecture research for materials property prediction, band gap prediction represents both a challenge and opportunity. Current evidence suggests that hybrid approaches—combining LLM-based data extraction with traditional machine learning—can achieve performance improvements (19% MAE reduction) over models trained on human-curated databases [58]. As foundation models continue to evolve in materials science, their ability to leverage multimodal data (text, structure, properties) may further transform band gap prediction by enabling more accurate, generalizable, and interpretable models that accelerate the discovery of novel materials with tailored electronic properties.
Beyond Baseline BERT: Optimization Strategies for Peak Performance
Tackling Data Sparsity with Advanced Positional Embeddings
In the field of AI-driven drug discovery, data sparsity presents a fundamental bottleneck. The chemical space is nearly infinite, while experimentally validated molecular property data is scarce, expensive to produce, and often limited to specific chemical regions. This sparsity challenge is particularly acute in materials property prediction research, where accurate predictions require models to generalize effectively from limited examples. The Bidirectional Encoder Representations from Transformers (BERT) architecture has emerged as a powerful framework for molecular property prediction, frequently processing molecular structures as Simplified Molecular Input Line Entry System (SMILES) strings. However, standard BERT implementations with basic positional embeddings struggle with the complex, non-sequential relationships inherent in molecular data and often fail to extrapolate to structures longer than those seen in training.
Advanced positional embeddings have recently surfaced as a critical solution to these limitations. By more effectively encoding the positional relationships between atoms and substructures in molecular representations, these advanced methods enable transformer models to better capture the intricate syntax of chemical "language," thereby improving generalization, enabling zero-shot learning for novel compounds, and ultimately tackling the core challenge of data sparsity in molecular sciences.
Positional Embedding Fundamentals in Transformer Architectures
Transformers, unlike their recurrent neural network predecessors, process all tokens in a sequence simultaneously through their self-attention mechanisms. This architectural strength creates a fundamental limitation: native transformers are permutation-invariant and cannot inherently discern the order of input tokens. Positional embeddings solve this problem by injecting information about token position into the model, allowing it to understand sequence ordering crucial for interpreting molecular structures.
The self-attention mechanism computes outputs as a weighted sum of values, where weights are based on compatibility between queries and keys: Attention(Q, K, V) = softmax(QK^T/√d_model)V [61]. Without positional information, rearranging input tokens would produce identical attention outputs regardless of order. Positional embeddings modify this mechanism to ensure the model recognizes that "CCN" represents a different molecule than "CNC" despite containing identical atoms.
Theoretical Foundation and Molecular Applications
In molecular property prediction, positional embeddings must capture more than simple sequence position; they must encode the complex topological relationships between atoms that define molecular structure and function. Traditional sequential embeddings often fail to capture these relationships, leading to inefficient learning and poor generalization on sparse molecular datasets. Advanced embeddings address this by modeling relative positions, rotational constraints, or two-dimensional spatial relationships that more closely mirror chemical reality.
Comparative Analysis of Positional Embedding Methods
Absolute Positional Embeddings
Absolute positional embeddings assign a unique vector to each position in the sequence using either predetermined sinusoidal functions or learnable parameters.
Table 1: Absolute Positional Embedding Characteristics
| Feature | Sinusoidal | Learned |
|---|---|---|
| Definition | Predefined using sine/cosine functions with varying frequencies | Parameters learned during training |
| Generalization | Theoretical extrapolation capability | Limited to trained sequence lengths |
| Molecular Application | Rare in modern molecular transformers | Foundational in early SMILES-based BERT |
| Key Limitation | Struggles to capture relative positioning | Poor generalization to longer sequences |
The original Transformer paper proposed sinusoidal functions: PE(pos, 2i) = sin(pos/10000^(2i/d_model)) and PE(pos, 2i+1) = cos(pos/10000^(2i/d_model)) where pos is position and i is the dimension [61]. This method theoretically helps models learn relative positions due to its linear properties, but in practice, transformers with absolute embeddings struggle to recognize that position 5 and 6 are similarly related as position 105 and 106.
Relative Positional Embeddings
Relative positional embeddings encode the distance between tokens rather than their absolute positions, directly modeling the pairwise relationships between atoms in a molecular sequence.
Table 2: Relative Positional Embedding Implementation Approaches
| Aspect | Additive Bias Method | Key-Query Integration |
|---|---|---|
| Mechanism | Adds learnable biases to attention scores based on relative distance | Incorporates relative position into key and query calculations |
| Computational Impact | Moderate increase in parameters | Higher computational overhead |
| Sequence Length Handling | Clipping beyond threshold K | Typically uses clipped relative distance |
| Molecular Advantage | Captures local atomic interactions | Better models long-range molecular dependencies |
Relative methods modify the attention calculation to incorporate pairwise distance: z_i = Σ_j softmax((x_iW_Q)(x_jW_K)^T + a_ij)/√d * x_jW_V where a_ij is a learnable bias term representing the relative position between token i and j [62]. This approach directly informs the model about the spatial relationships between atoms regardless of their absolute positions in the sequence.
Rotary Position Embedding (RoPE)
Rotary Position Embedding (RoPE) represents a breakthrough approach that encodes absolute position with a rotation operation while naturally incorporating relative position information in the attention mechanism.
Table 3: Rotary Position Embedding Analysis
| Characteristic | Description | Molecular Relevance |
|---|---|---|
| Core Mechanism | Rotates queries and keys using rotation matrices | Preserves relative position information regardless of sequence length |
| Extrapolation Capability | Strong performance on longer sequences | Critical for complex molecules exceeding training length |
| Computational Efficiency | No additional parameters; minimal overhead | Enables processing of large molecular libraries |
| Theoretical Foundation | Applies rotation transformation to token embeddings | Maintains geometric relationships between atomic representations |
RoPE transforms queries and keys using a rotation matrix: f(q, m) = R(m)q where R(m) is a rotation matrix that incorporates position m through angle θ [61]. For a pair of tokens at positions m and n, their dot product after rotation depends only on their relative distance (m-n), not their absolute positions. This property makes RoPE particularly effective for molecular sequences where the relationship between distant atoms determines key properties.
Figure 1: RoPE Integration in Transformer Workflow
Experimental Comparison in Molecular Property Prediction
Methodology for Evaluating Positional Embeddings
Recent studies have established rigorous protocols for evaluating positional embeddings in molecular BERT models. The standard approach follows a two-stage framework: pretraining on large unlabeled molecular datasets followed by fine-tuning on specific property prediction tasks.
Pretraining Phase: Models undergo masked language modeling pretraining on extensive SMILES datasets (e.g., 7.9 million instances) [63]. During this phase, 15% of tokens are randomly masked, and the model learns to predict them based on context. Different positional embeddings influence how effectively models learn molecular syntax and long-range dependencies.
Fine-tuning Phase: Pretrained models are adapted to specific prediction tasks (ADMET properties, bioactivity, toxicity) using labeled datasets. Performance is measured using domain-specific metrics: ROC-AUC for classification, RMSE for regression, with emphasis on zero-shot performance on novel molecular scaffolds.
Critical Experimental Considerations:
- Sequence Length Variability: Evaluation across molecules of varying lengths
- Scaffold Diversity: Assessment on structurally distinct compounds
- Data Efficiency: Performance with limited training examples
Quantitative Performance Comparison
Table 4: Experimental Results of Positional Embeddings on Molecular Tasks
| Embedding Type | Accuracy (%) | Sequence Length Extrapolation | Data Efficiency | Zero-Shot Performance |
|---|---|---|---|---|
| Absolute (Sinusoidal) | 84.3 | Poor (<15% beyond trained length) | Low (requires ~70% more data) | Limited (F1: 0.62) |
| Absolute (Learned) | 85.1 | Very Poor (fails beyond max length) | Medium | Limited (F1: 0.59) |
| Relative Key | 87.2 | Good (65% performance maintained) | Medium-High | Good (F1: 0.71) |
| RoPE | 88.7 | Excellent (82% performance maintained) | High | Strong (F1: 0.76) |
Recent research examining BERT for molecular-property prediction demonstrated that models with RoPE embeddings achieved superior accuracy (up to 88.7%) and significantly better generalization to longer sequences compared to absolute and relative baselines [63]. The rotary approach maintained 82% of its performance on sequences 50% longer than those seen during training, while absolute embeddings virtually failed under the same conditions.
Case Study: Positional Embeddings for COVID-19 Drug Discovery
The COVID-19 pandemic highlighted the critical need for models that could rapidly predict molecular properties for novel compounds. Researchers evaluated various positional embeddings on BERT models tasked with predicting antiviral activity against SARS-CoV-2 [63].
Figure 2: COVID-19 Antiviral Prediction Experimental Design
The study found that RoPE-based models significantly outperformed other embeddings, particularly in zero-shot scenarios involving structurally novel compounds. This advantage stemmed from RoPE's ability to maintain stable relative position relationships even for molecular sequences with unfamiliar scaffolds or longer chain lengths.
Implementation Guide: Advanced Embeddings for Molecular BERT
Research Reagent Solutions
Table 5: Essential Research Components for Positional Embedding Experiments
| Component | Function | Implementation Example |
|---|---|---|
| SMILES Tokenizer | Converts SMILES strings to token sequences | Byte-pair encoding adapted for chemical syntax |
| Positional Embedding Module | Injects position information into transformer | RoPE implementation as PyTorch module |
| Molecular Datasets | Benchmarks for pretraining and fine-tuning | COVID-19 bioassay, ADMET benchmarks |
| Evaluation Framework | Standardized assessment across tasks | Multi-task metrics with scaffold splitting |
Practical Integration of RoPE in Molecular BERT
Implementing RoPE requires modifying the attention mechanism to apply rotation matrices to queries and keys based on their positions:
This rotation preserves the relative position information through the dot product: q_rot(m) • k_rot(n) = R(m-n)(q • k), where the output depends only on the relative distance (m-n) [61].
The evolution of positional embeddings from absolute to rotary representations marks significant progress in tackling data sparsity for molecular property prediction. RoPE's mathematical elegance and empirical superiority make it particularly well-suited for molecular BERT applications, where capturing precise relationships between distant atomic constituents often determines prediction accuracy.
As molecular property prediction advances, several research directions emerge: (1) developing domain-adapted positional embeddings that incorporate chemical knowledge beyond simple sequence position; (2) creating dynamic embedding strategies that adjust to molecular graph topology rather than linear sequences; and (3) designing multi-modal embeddings that simultaneously capture sequence, graph, and spatial relationships in molecular data.
For researchers and drug development professionals, embracing advanced positional embeddings like RoPE can substantially enhance model performance on sparse data regimes common in early-stage discovery. These technical improvements translate to more accurate prediction of ADMET properties, bioactivity, and toxicity for novel compounds, ultimately accelerating the drug discovery pipeline and reducing experimental costs.
Integrating Active Learning for Efficient Experimental Design
The application of BERT architecture in materials property prediction represents a paradigm shift in computational drug development and materials informatics. This approach integrates transformer-based deep learning with strategic experimental design to significantly accelerate the discovery pipeline. Active learning (AL), a semi-supervised machine learning approach that iteratively selects the most informative data points for labeling, has emerged as a critical component for optimizing resource allocation in experimental sciences [9]. When combined with BERT's ability to generate rich molecular representations from unlabeled data, this synergy creates a powerful framework for efficient experimental design. The integration addresses a fundamental challenge in pharmaceutical research: the prohibitive cost and time requirements of exhaustive experimental testing. By prioritizing compounds with the highest potential, researchers can focus resources on the most promising candidates, dramatically improving the efficiency of drug discovery workflows [9] [24].
Comparative Analysis of BERT-Enhanced Active Learning Approaches
Performance Benchmarking Across Methodologies
Table 1: Quantitative performance comparison of predictive modeling approaches
| Methodology | Application Domain | Dataset | Key Performance Metric | Performance Result | Comparative Advantage |
|---|---|---|---|---|---|
| BERT + Bayesian AL [9] | Molecular Toxicology | Tox21 & ClinTox | Iteration Reduction | 50% fewer iterations | Equivalent toxic compound identification with half the experimental cycles |
| Pretrained BERT [9] | Molecular Representation | 1.26M compounds | Embedding Quality | Structured embedding space | Reliable uncertainty estimation with limited labeled data |
| Deep Transfer Learning [64] | Formation Energy Prediction | Experimental Hold-out Set | Mean Absolute Error | 0.064 eV/atom | Outperforms DFT computations (>0.076 eV/atom) |
| BERT Pathology Model [65] | Medical Text Analysis | Bone Marrow Synopses | Micro-average F1 Score | 0.779 ± 0.025 | Effective semantic label mapping with minimal training data |
| Traditional DFT [66] | Formation Energy Prediction | Multiple Databases | Mean Absolute Error | 0.078-0.095 eV/atom | Baseline for AI-based improvement |
| ElemNet [66] | Formation Energy Prediction | Experimental Dataset | Mean Absolute Error | ~0.15 eV/atom | Improved through transfer learning (~0.06 eV/atom) |
Integration Advantages in Experimental Design
The comparative data reveals that BERT-enhanced active learning systems consistently outperform traditional computational approaches across multiple domains. In molecular property prediction, the integration of pretrained BERT with Bayesian active learning achieves equivalent performance to conventional methods with 50% fewer experimental iterations [9] [24]. This efficiency gain stems from BERT's ability to create structured embedding spaces from extensive unlabeled molecular data (1.26 million compounds), enabling reliable uncertainty estimation even when labeled data is scarce [9].
In materials science, deep transfer learning approaches demonstrate similar advantages, with AI models predicting formation energy from materials structure and composition with significantly better accuracy than Density Functional Theory (DFT) computations themselves [64]. This breakthrough is particularly notable as it surmounts the inherent discrepancies between DFT computations and experimental observations that have traditionally limited predictive modeling in materials science [66].
For specialized domains like pathology, BERT-based active learning enables effective information extraction from complex medical texts with minimal training data, achieving robust performance (F1 score: 0.779) with only 500 labeled examples developed through an iterative active learning process [65].
Experimental Protocols and Methodologies
BERT-Enhanced Bayesian Active Learning Framework
Experimental Protocol for Molecular Property Prediction:
The integrated BERT and Bayesian active learning methodology follows a structured workflow [9]:
Pretraining Phase: A transformer-based BERT model (MolBERT) is initially pretrained on 1.26 million unlabeled compounds to learn general molecular representations without task-specific labels. This pretraining captures fundamental chemical patterns and relationships within a broad chemical space.
Initial Model Setup: A small, balanced initial labeled set is created (e.g., 100 molecules with equal positive/negative representation) through random selection from the available training data. Scaffold splitting with 80:20 ratio ensures distinct training and testing sets that do not share core structural motifs, providing better generalization assessment [9].
Bayesian Active Learning Cycle:
- Model Training: The pretrained BERT model is fine-tuned on the current labeled set, leveraging transfer learning to adapt general molecular representations to the specific prediction task.
- Uncertainty Estimation: Bayesian methods quantify prediction uncertainties for all unlabeled compounds in the pool, capturing both epistemic (model) and aleatoric (data) uncertainty.
- Acquisition Function: Bayesian Active Learning by Disagreement (BALD) selects the most informative samples by maximizing the expected information gain about model parameters [9]. BALD computes the mutual information between model parameters and predictions, identifying samples where model uncertainty is highest.
- Experimental Labeling: The selected compounds undergo experimental testing (e.g., toxicity assays) to obtain ground-truth labels.
- Dataset Expansion: Newly labeled compounds are added to the training set, and the cycle repeats for predetermined iterations or until performance targets are met.
Performance Evaluation: The method is validated on benchmark datasets like Tox21 (≈8,000 compounds across 12 toxicity pathways) and ClinTox (1,484 compounds comparing FDA-approved and failed drugs), with metrics including early identification efficiency and calibration error [9].
Deep Transfer Learning for Materials Property Prediction
Experimental Protocol for Formation Energy Prediction [64] [66]:
Data Preparation: Utilize multiple DFT-computed databases (OQMD, Materials Project, JARVIS) containing formation energies for thousands of materials, alongside experimental datasets (e.g., SSUB database with 1,963 formation energies at 298.15K).
Source Model Training: Train a deep neural network (e.g., ElemNet or IRNet) on large DFT-computed source domains (e.g., ~341,000 materials in OQMD) to learn rich feature representations from materials structure and composition.
Transfer Learning Fine-tuning: Adapt the pretrained model to experimental observations through additional training on smaller, accurate experimental datasets. This fine-tuning process adjusts model parameters to bridge the discrepancy between DFT computations and experimental values.
Validation: Evaluate the model on hold-out experimental test sets (e.g., 137 entries) comparing performance against pure DFT computations and models trained from scratch on experimental data only.
Active Learning for Pathology Text Analysis
Experimental Protocol for Semantic Label Generation [65]:
Iterative Label Development: Employ active learning to develop a comprehensive set of semantic labels for bone marrow aspirate pathology synopses through 9 iterative cycles, expanding from 10 to 21 labels and 50 to 500 samples.
BERT Model Training: Fine-tune a BERT model pretrained on general domain text (800 million words) using the labeled pathology synopses, leveraging the transformer's attention mechanisms to capture syntactic and semantic relationships in medical text.
Embedding Extraction: Extract classification (CLS) feature vectors from the final BERT layer, representing embeddings that capture diagnostically relevant semantic information from pathology text.
Multi-label Classification: Map the extracted embeddings to one or more semantic labels representing diagnostic categories, using the model to automatically annotate pathology synopses with clinically relevant concepts.
Workflow Visualization
Figure 1: BERT-enhanced Bayesian active learning workflow for molecular property prediction, illustrating the cyclic interaction between computational modeling and experimental validation. [9] [24]
Table 2: Key research reagents and computational resources for BERT-based active learning experiments
| Resource | Type | Specification | Research Application |
|---|---|---|---|
| Tox21 Dataset [9] | Biological Assay Data | ≈8,000 compounds, 12 toxicity pathways, binary labels | Benchmark for molecular toxicology prediction models |
| ClinTox Dataset [9] | Clinical Trial Data | 1,484 compounds (FDA-approved vs failed drugs) | Comparison of drug safety profiles in clinical trials |
| OQMD Database [64] [66] | Computational Materials Data | ~341,000 materials with DFT-computed properties | Source domain for transfer learning of formation energy |
| Materials Project [64] [66] | Computational Materials Data | 30,000+ inorganic compounds with properties | Training and validation of materials property predictors |
| JARVIS Database [64] [66] | Computational Materials Data | 11,050 stable materials with formation energies | Comparative analysis of DFT computation accuracy |
| SSUB Database [66] | Experimental Materials Data | 1,963 formation energies at 298.15K | Ground truth validation for formation energy prediction |
| MolBERT Model [9] | Computational Algorithm | Transformer architecture pretrained on 1.26M compounds | Molecular representation learning for chemical space |
| BERT Base Model [65] | Natural Language Algorithm | Transformer trained on 800M words, medical domain adaption | Semantic information extraction from pathology text |
| BALD Acquisition [9] | Computational Method | Bayesian Active Learning by Disagreement | Optimal sample selection for experimental labeling |
The integration of BERT architectures with active learning frameworks represents a transformative advancement in experimental design for materials and drug discovery. The comparative data demonstrates that this approach consistently outperforms traditional computational methods, reducing experimental iterations by 50% in molecular toxicology prediction [9] and achieving superior accuracy to DFT in formation energy prediction [64]. The fundamental advantage stems from the synergy between BERT's ability to learn rich representations from unlabeled data and active learning's strategic selection of informative samples for experimental testing. This paradigm effectively bridges the gap between computational prediction and experimental validation, enabling researchers to navigate complex chemical and materials spaces with unprecedented efficiency. As these methodologies continue to evolve, they promise to significantly accelerate the discovery and development of novel therapeutic compounds and advanced materials.
The field of materials property prediction is undergoing a significant transformation, driven by the convergence of artificial intelligence and quantum computing. Within this context, the fusion of powerful classical language models like Bidirectional Encoder Representations from Transformers (BERT) with emerging Quantum Neural Networks (QNNs) represents a frontier of research with the potential to redefine computational efficiency and predictive accuracy. These architectural hybrids are being developed to tackle fundamental challenges in materials informatics and drug discovery, including data sparsity, high computational costs, and the need to model complex quantum mechanical interactions. This guide provides an objective comparison of emerging BERT-QNN architectures, detailing their performance against classical alternatives, underlying methodologies, and practical implementation considerations for researchers and drug development professionals.
Performance Comparison of BERT-QNN Hybrids
Experimental results from recent studies demonstrate that hybrid BERT-QNN models can match or exceed the performance of classical models while achieving significant gains in parameter efficiency. The following table summarizes key quantitative comparisons.
Table 1: Performance Comparison of BERT-QNN Hybrid Models vs. Classical Alternatives
| Model Name | Application Domain | Performance Metrics vs. Classical Baseline | Parameter Efficiency | Key Advantage |
|---|---|---|---|---|
| QFFN-BERT [67] | Natural Language Processing | Achieved up to 102.0% of the baseline BERT accuracy on SST-2 and DBpedia benchmarks [67]. | >99% reduction in parameters in the replaced Feedforward Network (FFN) modules [67]. | Superior data efficiency in few-shot learning scenarios. |
| PolyQT [68] | Polymer Property Prediction | R² values for ionization energy, dielectric constant, and glass transition temperature reached 0.85, 0.77, and 0.85, respectively, surpassing all classical benchmark models (GP, NN, RF, LSTM, Transformer) [68]. | Not explicitly quantified, but the model demonstrated superior performance under high data sparsity (40-80% sparsity levels) [68]. | Effectively addresses data sparsity issues; maintains high accuracy with limited data. |
| Quantum-Embedded GNN (QEGNN) [69] | Molecular Property Prediction | Consistently achieved higher accuracy and improved stability on multiple benchmark datasets [69]. | Significantly reduced parameter complexity cited as a hallmark of quantum advantage [69]. | Stable performance on current noisy quantum hardware ("Wukong" processor). |
| Hybrid Quantum Neural Network [70] | Entity Matching (NLP) | Reached similar performance as classical approaches (TF-IDF, neural networks) [70]. | Required an order of magnitude fewer parameters than its classical counterpart [70]. | Model trained on a quantum simulator is transferable to real quantum computers. |
Detailed Experimental Protocols and Methodologies
The performance gains outlined above are underpinned by specific architectural choices and training methodologies. This section details the experimental protocols for two primary hybrid approaches: replacing core BERT components with quantum circuits, and using BERT for initial feature extraction before a quantum classifier.
QFFN-BERT: Replacing the Feedforward Network
The QFFN-BERT architecture is a direct hybrid where the classical Feedforward Network (FFN) in a compact BERT variant is replaced with a Parameterized Quantum Circuit (PQC). This design is motivated by the fact that FFNs account for approximately two-thirds of the parameters in a standard Transformer encoder block [67].
Protocol:
- Circuit Design: The PQC incorporates several key features to ensure trainability and expressibility:
- A residual connection to stabilize training.
- Both
RYandRZrotation gates for increased expressibility. - An alternating entanglement strategy to create qubit correlations.
- Systematic variation of the PQC depth is performed to find a sweet spot between expressibility and the onset of barren plateau phenomena (vanishing gradients) [67].
- Integration: The quantum component is implemented using Qiskit and integrated into a PyTorch framework via TorchConnector [67].
- Training & Evaluation: The model is trained and evaluated on a classical simulator using standard NLP benchmarks like SST-2 (sentiment analysis) and DBpedia (topic classification). Performance is compared against a classical BERT baseline with an equivalent number of attention layers and hidden dimensions [67].
PolyQT and Quantum-Embedded Models: Feature Extraction and Classification
Another common protocol uses a classical BERT model for initial feature extraction, the output of which is then processed by a separate QNN for property prediction. This is prevalent in scientific domains like polymer and molecular property prediction.
Protocol:
- Data Representation: Polymer or molecular structures are represented as text strings (e.g., SMILES, DeepSMILES, Big-SMILES) [71] [68]. A tokenizer processes these strings into tokens suitable for the model.
- Feature Extraction: A pre-trained BERT model processes the tokenized sequences. The output from BERT's encoder (often the [CLS] token embedding or mean of token embeddings) serves as a rich, contextual feature vector representing the input structure [71] [9].
- Quantum Classification:
- The classical feature vector is mapped into a quantum state using an embedding circuit (e.g., angle embedding based on feature vector values) [68].
- A parametrized quantum circuit (PQC), or QNN, processes the embedded state. The structure of this PQC (e.g., number of qubits, arrangement of rotation and entanglement layers) is a critical hyperparameter [68].
- Measurements from the quantum circuit are used to generate the final prediction (e.g., regression value for a property, or a classification label).
- Training: The hybrid model is typically trained in a hybrid quantum-classical loop:
- The quantum circuit's measurements provide the loss value.
- A classical optimizer (e.g., Adam, SGD) computes gradients and updates both the classical (BERT) and quantum (PQC) parameters [70].
The workflow for this hybrid feature extraction and classification approach is visualized below.
The Scientist's Toolkit: Essential Research Reagents and Materials
Implementing BERT-QNN hybrids requires a suite of software tools and hardware access. The following table details the key components.
Table 2: Essential Research Tools for BERT-QNN Hybrid Model Development
| Tool Name | Type | Function in the Workflow |
|---|---|---|
| PyTorch / TensorFlow [67] | Classical ML Framework | Provides the foundational infrastructure for building, training, and managing the classical components of the model (e.g., the BERT model itself, classical embedding layers). |
| Hugging Face Transformers | Library | Offers easy access to pre-trained BERT models and tokenizers, significantly accelerating the feature extraction development phase [71] [9]. |
| Qiskit [67] [70] | Quantum Computing SDK | (IBM) Allows for the design and simulation of parameterized quantum circuits (PQCs). Includes TorchConnector for seamless integration with PyTorch, enabling gradient propagation [67]. |
| Cirq [70] | Quantum Computing SDK | (Google) A Python library for writing, manipulating, and optimizing quantum circuits and running them on simulators and real quantum computers. |
| Lambeq [72] | QNLP Toolkit | A specialized Python toolkit for Quantum Natural Language Processing (QNLP), which converts sentences into quantum circuits following the DisCoCat model, facilitating semantic tasks. |
| Quantum Simulator | Computational Resource | A classical software simulator of a quantum computer (e.g., Qiskit Aer). Essential for algorithm development, debugging, and initial training runs before deploying on expensive quantum hardware [67] [70]. |
| NISQ Computer | Hardware | Noisy Intermediate-Scale Quantum computers (e.g., IBM's cloud-based quantum systems). Required for final validation and testing on real, albeit noisy, quantum devices [70] [69]. |
Critical Analysis and Research Outlook
While the experimental data is promising, several challenges and limitations define the current research frontier. A primary constraint is the reliance on Noisy Intermediate-Scale Quantum (NISQ) hardware, which is characterized by limited qubit counts, short coherence times, and high error rates [73]. This currently restricts the complexity of feasible quantum circuits and the size of problems that can be tackled. Furthermore, researchers must carefully navigate the expressibility-trainability trade-off; while increasing quantum circuit depth can enhance representational power, it also increases susceptibility to the barren plateau problem, where gradients vanish and training becomes impossible [67].
Future research is directed towards overcoming these hurdles. The development of more sophisticated error mitigation techniques and the eventual arrival of fault-tolerant quantum hardware will be pivotal [73]. There is also a strong focus on hybrid quantum-classical algorithms that more intelligently divide labor between classical and quantum components to maximize the strengths of each paradigm [73] [68]. As these technologies mature, BERT-QNN hybrids are poised to make significant impacts in areas like personalized medicine through patient-specific molecular simulations and the efficient exploration of vast chemical spaces for de novo drug design [74] [73].
The application of BERT architecture to materials property prediction represents a significant advancement in computational drug discovery and materials science. However, the inherent "black-box" nature of complex deep learning models poses a significant challenge for research and development professionals who require transparent, interpretable predictions for critical decision-making. This guide provides a comprehensive comparison of the two predominant approaches for enhancing model interpretability: game-theoretic methods such as SHapley Additive exPlanations (SHAP) and attention weight visualization techniques exemplified by tools like BertViz. Within the context of molecular property prediction, these interpretability frameworks serve complementary roles—game-theoretic approaches quantify feature importance post-hoc, while attention visualization provides intrinsic insights into model reasoning by illuminating the internal computational processes of transformer architectures.
Comparative Analysis of Interpretability Approaches
The table below summarizes the core characteristics, strengths, and limitations of game-theoretic and attention-based interpretability methods.
Table 1: Comparison of Interpretability Approaches for BERT in Property Prediction
| Feature | Game-Theoretic Approaches (e.g., SHAP) | Attention Weight Visualization (e.g., BertViz) |
|---|---|---|
| Core Principle | Computes feature importance based on cooperative game theory, quantifying each feature's marginal contribution to the prediction [75]. | Visualizes the attention mechanism within transformer models, showing how input tokens weigh each other when producing representations [76] [77]. |
| Interpretability Type | Post-hoc, model-agnostic explanation [75]. | Primarily intrinsic and model-specific [76]. |
| Typical Output | Feature importance scores and summary plots (e.g., beeswarm plots) [75]. | Interactive visualizations of attention flows (e.g., head view, model view) [76] [77]. |
| Key Strength | Provides a mathematically grounded, quantitative measure of feature contribution; works with any model [75]. | Offers a direct, intuitive view into the model's "reasoning process" during computation [76] [78]. |
| Primary Limitation | Computationally expensive; explanations are approximations separate from the model's actual inner workings [75]. | The relationship between attention weights and model output is not always straightforward; may not be the sole source of model behavior [76]. |
| Application Example | Explaining which molecular descriptors (e.g., molecular weight, lipophilicity) most influenced a toxicity prediction [75] [3]. | Visualizing how a BERT model attends to different atoms in a SMILES string when predicting a molecular property like solubility [76] [63]. |
Experimental Protocols and Performance Data
Implementation of Attention Visualization with BertViz
BertViz is an open-source tool that visualizes the attention mechanism in transformer models at multiple levels of granularity—model-wide, attention head-level, and neuron-level [76] [77]. The following workflow details its standard implementation for analyzing a molecular property prediction model.
Experimental Protocol 1: Visualizing Attention in SMILES-Based BERT
- Model and Tokenization: Load a pre-trained or fine-tuned BERT model configured to return attention weights. The input is a SMILES (Simplified Molecular Input Line Entry System) string representing a molecular structure. A tokenizer converts this string into token IDs and subword tokens [77] [63].
- Model Inference: Pass the tokenized input through the model to obtain both the prediction (e.g., solubility, toxicity) and the attention tensors [77].
- Visualization: Use BertViz's API to render interactive visualizations. Common views include:
- Head View: Shows how attention flows between tokens for one or more attention heads in a single layer, revealing patterns like fixation on specific functional groups [76] [77] [78].
- Model View: Provides a bird's-eye view of attention across all layers and heads, helping identify high-level data flow patterns [77].
- Neuron View: Visualizes the individual query, key, and value vectors that compute attention, showing how specific attention patterns are formed [76] [77].
The following diagram illustrates this experimental workflow.
Implementation of Game-Theoretic Explanations with SHAP
SHAP is a prominent game-theoretic approach that explains a model's output by calculating the marginal contribution of each feature to the prediction across all possible feature combinations [75]. The protocol below applies to explaining a molecular property predictor.
Experimental Protocol 2: Explaining Predictions using SHAP
- Model and Background Data: Select a trained model (can be a BERT model whose outputs are used as features, or a separate predictor) and a representative background dataset (e.g., a random sample of molecules) to establish a baseline for expected predictions [75].
- Explanation Generation: For a given molecule to be explained (the "instance"), the SHAP library computes Shapley values. This involves creating many perturbed versions of the instance by masking subsets of its features (e.g., atoms, bonds, or molecular descriptors) and using the background dataset to fill them in. The model's prediction on these perturbed inputs is used to compute the average marginal contribution of each feature [75].
- Visualization and Analysis: The computed Shapley values are visualized to show which features pushed the model's prediction higher (positive contribution) or lower (negative contribution) relative to the baseline prediction. Common plots include force plots for single-instance explanations and summary plots for dataset-level analysis [75].
Quantitative Performance Comparison
Experimental studies have demonstrated the utility of both interpretability approaches in real-world molecular prediction tasks. The following table summarizes key performance metrics from recent research.
Table 2: Experimental Performance in Molecular Property Prediction
| Model / Interpretability Method | Dataset | Task | Key Metric | Result & Interpretation Insight |
|---|---|---|---|---|
| Pretrained BERT with Active Learning [3] | Tox21, ClinTox | Toxicity Identification | Equivalent identification accuracy with 50% fewer iterations than conventional AL. | SHAP-like analysis revealed that pretrained BERT representations created a structured embedding space, enabling more reliable uncertainty estimation for sample acquisition [3]. |
| SMG-BERT (Integrates 3D geometry) [79] | 12 Benchmark Molecular Datasets | Property Prediction | Consistently outperformed existing state-of-the-art models. | Attention visualization enabled interpretability consistent with chemical logic, highlighting relevant substructures and stereochemistry due to integrated NMR and bond energy features [79]. |
| Geometry-based BERT (GEO-BERT) [4] | DYRK1A Inhibitor Screening | Prospective Validation | Identified two potent novel inhibitors (IC50: <1 μM). | Attention mechanisms, guided by 3D positional relationships (atom-atom, bond-bond, atom-bond), likely helped the model focus on spatially relevant structural motifs for activity [4]. |
The Scientist's Toolkit: Essential Research Reagents
The following table details key computational tools and resources essential for implementing the interpretability methods discussed in this guide.
Table 3: Key Research Reagents and Computational Tools
| Item Name | Function / Role | Specification / Note |
|---|---|---|
| BertViz [76] [77] | An interactive visualization tool for rendering attention mechanisms in transformer models directly within Jupyter or Colab notebooks. | Supports most HuggingFace models (BERT, GPT-2, T5, etc.). Provides Head, Model, and Neuron views. |
| SHAP Library [75] | A Python library that computes Shapley values from game theory to explain the output of any machine learning model. | Model-agnostic. Offers various explainers (e.g., KernelExplainer, DeepExplainer) for different model types. |
| HuggingFace Transformers [77] | A Python library providing thousands of pre-trained transformer models for a wide range of tasks. | Essential for loading and running models compatible with BertViz. Simplifies model fine-tuning on custom datasets. |
| RDKit | An open-source cheminformatics toolkit used for processing SMILES strings, generating molecular fingerprints, and calculating molecular descriptors. | Often used for preprocessing molecular inputs for BERT models and for post-hoc analysis of feature importance [79]. |
| SMILES Strings [63] [80] | A line notation for representing molecular structures as text, serving as the primary input sequence for molecular BERT models. | Must be tokenized (e.g., via Byte-Pair Encoding) before being fed into a transformer model. |
| Molecular Datasets (e.g., Tox21, ClinTox) [3] | Curated public datasets used for training and benchmarking molecular property prediction models. | Typically contain SMILES strings and associated experimental property/activity labels. |
Within BERT-based architectures for materials property prediction, the initial step of converting raw molecular structures into model-processable tokens is a critical determinant of performance. Tokenization strategies for Simplified Molecular Input Line Entry System (SMILES) strings directly influence a model's ability to learn meaningful chemical representations, impacting predictive accuracy across diverse pharmaceutical applications. This guide provides an objective comparison of contemporary tokenization methodologies, evaluating their experimental performance and implementation considerations for drug discovery researchers.
The Tokenization Landscape for Molecular Representations
Molecular tokenization serves as the foundational bridge between chemical structures and machine learning models. Unlike natural language processing, where tokens typically represent words or sub-words, chemical tokenization decomposes molecular string representations into constituent units that preserve structural meaning while facilitating efficient model training.
Core Tokenization Philosophies
Two primary philosophies dominate molecular tokenization approaches for BERT-based architectures:
- Chemistry-Agnostic Tokenization: Treats SMILES as generic text strings, applying standard NLP tokenization methods like Byte Pair Encoding (BPE) to allow the model to learn chemical relationships from data patterns alone. This approach requires substantial training data but offers broad generalizability. [81]
- Chemistry-Aware Tokenization: Incorporates domain knowledge through preprocessing that identifies chemically meaningful substructures before model training. This approach improves data efficiency but requires specialized chemical expertise in implementation. [81]
Comparative Analysis of Tokenization Methods
Fundamental SMILES Tokenization Approaches
| Method | Core Principle | Key Advantages | Key Limitations | Representative Performance |
|---|---|---|---|---|
| Atom-wise SMILES | Character-level tokenization of SMILES strings | Simple implementation; Widely supported | Limited token diversity; Fails to capture chemical context | Baseline performance on MoleculeNet benchmarks [82] |
| Byte Pair Encoding (BPE) | Merges frequent character pairs into tokens | Reduces vocabulary size; Captures common substrings | May create chemically meaningless tokens | Competitive with graph networks on 6/8 MoleculeNet tasks [83] [81] |
| SMILES Pair Encoding (SmilesPE) | SMILES-optimized BPE variant | Improved chemical relevance over standard BPE | Limited contextual awareness | Moderate improvements over atom-wise tokenization [82] |
Advanced Chemistry-Aware Tokenization Strategies
| Method | Core Principle | Vocabulary Characteristics | Performance Highlights | Data Efficiency |
|---|---|---|---|---|
| Atom-in-SMILES (AIS) | Replaces atomic symbols with environment-aware tokens | 10x token diversity increase over SMILES; Lower repetition rates (10% reduction) [82] | Superior in regression/classification tasks; 7% improvement in binding affinity prediction [84] | High - achieves strong results with standard dataset sizes |
| MolBERT Morgan Fingerprints | Uses circular atom-centered substructures as tokens | 13K chemically meaningful tokens [81] | 83.9% ROC-AUC on Tox21; 2-4% improvements over graph methods [81] | Excellent - effective with only 4M training compounds [81] |
| Hybrid Fragment-SMILES | Combines high-frequency fragments with atomic tokens | Balanced vocabulary; Mitigates token frequency imbalance [85] | Enhanced ADMET prediction; Optimal with 100-150 fragment tokens [85] [84] | High - outperforms SMILES tokenization in multi-task learning |
| SELFIES with BPE | Robust molecular representation guaranteeing validity | Similar atom/bond diversity to SMILES | Competitive but not superior to optimized SMILES approaches [83] | Moderate - requires standard dataset sizes |
Quantitative Performance Comparison Across Benchmarks
| Tokenization Method | Tox21 (ROC-AUC) | ClinTox (ROC-AUC) | HIV (ROC-AUC) | BBB Penetration (ROC-AUC) | Training Data Scale |
|---|---|---|---|---|---|
| Atom-wise SMILES | 0.791 [9] | 0.824 [9] | 0.763 [83] | 0.842 [83] | 77M compounds for optimal performance [81] |
| BPE with SMILES | 0.801 [83] | 0.831 [83] | 0.769 [83] | 0.851 [83] | 77M compounds for optimal performance [81] |
| MolBERT Morgan | 0.839 [81] | 0.858 [81] | - | - | 4M compounds [81] |
| AIS Tokenization | +3-5% over baseline [82] | +3-5% over baseline [82] | +3-5% over baseline [82] | +3-5% over baseline [82] | Standard dataset sizes |
| Atom Pair Encoding (APE) | 0.821 [83] | 0.847 [83] | 0.781 [83] | 0.863 [83] | Standard dataset sizes |
Experimental Protocols and Methodologies
Standardized Evaluation Framework
To ensure fair comparison across tokenization strategies, researchers have established consistent experimental protocols:
Dataset Preparation and Splitting
- Scaffold Splitting: Partitions molecules according to Bemis-Murcko scaffold representations using 80:20 train-test ratios, ensuring distinct structural motifs between sets to evaluate generalization capability. [9]
- Initial/Pool Set Construction: For active learning experiments, creates balanced initial sets (e.g., 100 molecules with equal positive/negative representation) with remaining training data forming the pool set. [9]
Performance Metrics
- ROC-AUC: Primary metric for binary classification tasks (e.g., toxicity, activity).
- Expected Calibration Error: Measures reliability of uncertainty estimates in Bayesian active learning. [9]
- Token Repetition Rate: Quantifies token diversity and potential degeneration issues. [82]
Implementation Methodologies
Chemistry-Agnostic Implementation (ChemBERTa)
- Pretraining: Masked language modeling on 77M compound datasets.
- Architecture: RoBERTa-base with 6 layers, 12 attention heads.
- Tokenization: BPE with vocabulary sizes ranging from 591-52K tokens.
- Fine-tuning: Task-specific heads for molecular property prediction. [81]
Chemistry-Aware Implementation (MolBERT)
- Token Generation: Morgan fingerprints with radius 1 (atom plus immediate neighbors).
- Vocabulary Construction: 13,325 unique substructure identifiers from 4M training molecules.
- Multi-Task Pretraining: Simultaneous prediction of 200 molecular properties.
- Architecture: BERT-base with optimized chemical token embeddings. [81]
Hybrid Tokenization Workflow
- Fragment Library Construction: High-frequency molecular fragments from chemical databases.
- Frequency Analysis: Fragment occurrence statistics to determine optimal cutoff (typically 100-150 tokens).
- Vocabulary Integration: Combined fragment and atomic tokens with frequency-based weighting.
- Model Training: Transformer architecture with multi-task learning objectives. [85]
Visualizing Tokenization Workflows and Relationships
Molecular Tokenization Strategy Decision Framework
Atom-in-SMILES (AIS) Tokenization Process
The Scientist's Toolkit: Essential Research Reagents
| Resource Category | Specific Resource | Function in Tokenization Research | Implementation Notes |
|---|---|---|---|
| Benchmark Datasets | Tox21 (≈8,000 compounds, 12 toxicity pathways) [9] | Standardized evaluation of toxicity prediction models | 6.24% active compounds; Address class imbalance |
| Benchmark Datasets | ClinTox (1,484 FDA-approved/failed drugs) [9] | Binary classification of clinical trial toxicity | Combined FDA-approved and failed trial compounds |
| Benchmark Datasets | ZINC Database (1.26M+ compounds) [9] [84] | Pretraining and token vocabulary construction | Source for frequency-based token selection |
| Software Libraries | Hugging Face Transformers [83] | BERT model implementation and training | Extensive pretrained model repository |
| Software Libraries | RDKit [81] | Molecular processing and descriptor calculation | Used for MTR pretraining in ChemBERTa-2 |
| Software Libraries | Chemprop [83] | Graph neural network baseline comparisons | Established benchmark for molecular property prediction |
| Evaluation Metrics | ROC-AUC [9] [83] | Primary performance metric for classification | Standard across molecular prediction tasks |
| Evaluation Metrics | Expected Calibration Error [9] | Uncertainty quantification for active learning | Critical for Bayesian experimental design |
| Evaluation Metrics | Token Repetition Rate [82] | Measures tokenization scheme degeneration | Lower values indicate more informative tokenization |
Tokenization strategy selection presents fundamental trade-offs between data efficiency, implementation complexity, and predictive performance. Chemistry-agnostic approaches (BPE, atom-wise) provide solid baselines and benefit from extensive data availability, while chemistry-aware methods (AIS, Morgan fingerprints, hybrid approaches) offer superior sample efficiency and performance for specialized applications. For BERT-based materials property prediction, emerging hybrid tokenization strategies that balance atomic and fragment-level information demonstrate particular promise for ADMET optimization and active learning applications. The optimal approach depends on specific research constraints, including dataset scale, computational resources, and target application domains.
Proof in Performance: Validating BERT Against State-of-the-Art Models
The accurate prediction of molecular and material properties is a cornerstone of modern drug development and materials science. Among the various machine learning architectures applied to this challenge, transformer-based models, particularly those derived from the BERT architecture, have emerged as a powerful approach. This guide provides an objective comparison of performance across three standard benchmarks—Tox21, ClinTox, and MatBench—focus on BERT-based models and their alternatives. The evaluation encompasses traditional machine learning methods, graph neural networks, and the latest transformer-based architectures, providing researchers with a comprehensive overview of the current landscape to inform model selection and development.
A critical issue in benchmarking, particularly for the Tox21 dataset, is benchmark drift. The original Tox21 Data Challenge dataset has been altered in popular benchmarks like MoleculeNet and OGB, with changes including removed molecules, redesigned data splits, and imputed missing labels, rendering cross-study comparisons unreliable [86]. This analysis prioritizes results from the recently established reproducible Tox21 leaderboard that restores the original challenge conditions to ensure valid comparisons [86].
Benchmarking Datasets and Experimental Protocols
Dataset Specifications and Evaluation Metrics
The benchmarks covered in this guide represent diverse challenges in molecular and materials informatics, from toxicity prediction to material property forecasting.
Table 1: Benchmark Dataset Specifications
| Dataset | Primary Task | Samples | Endpoints/Properties | Key Metrics | Data Splitting |
|---|---|---|---|---|---|
| Tox21 | Toxicity prediction | 12,060 training, 647 test [87] | 12 binary toxicity assays [86] | Mean AUC (Area Under ROC Curve) [86] | Original challenge split [86] |
| ClinTox | Clinical toxicity | 1,484 compounds [3] | 2 tasks: FDA approval status and clinical trial toxicity failure [88] | AUC, Balanced Accuracy [88] | Scaffold split (80:20) [3] |
| MatBench | Materials property prediction | 312 to 132,752 across 13 tasks [43] | Optical, thermal, electronic, thermodynamic, tensile, elastic properties [43] | MAE, RMSE, R² | Nested cross-validation [43] |
Standardized Experimental Protocols
Consistent experimental protocols are essential for fair model comparison:
Tox21 Protocol: The reproducible leaderboard implements the original challenge protocol [86]. Models are trained on the original 12,060 compounds and evaluated on the held-out test set of 647 compounds. Predictions are generated via a standardized API that accepts SMILES strings and returns probabilities for all 12 endpoints. The primary metric is the mean AUC across all endpoints [86].
ClinTox Protocol: Studies typically employ scaffold splitting to separate training and test sets based on core molecular structures, ensuring evaluation of generalization to novel chemotypes [3]. For active learning experiments, initial sets of 100 molecules are randomly selected with balanced class representation, with models iteratively selecting additional informative samples from a pool set [3].
MatBench Protocol: The benchmark uses a consistent nested cross-validation procedure for error estimation across all 13 tasks [43]. This approach mitigates model and sample selection biases by maintaining separate validation sets within training folds for hyperparameter tuning, with final performance reported on held-out test sets.
Performance Comparison of Modeling Approaches
Quantitative Performance on Tox21 and ClinTox
Table 2: Performance Comparison on Tox21 and ClinTox Benchmarks
| Model Architecture | Tox21 (Mean AUC) | ClinTox (AUC) | Key Features |
|---|---|---|---|
| DeepTox (2015) | 0.831 [86] | - | Original Tox21 winner; ensemble-based deep learning |
| Self-Normalizing NN (SNN) | Competitive with DeepTox [86] | - | Descriptor-based neural network |
| Random Forest | Outperformed XGBoost [86] | - | Traditional ensemble method |
| Chemprop | Evaluated [86] | - | Message passing neural network |
| BERT-based (ChemLM) | Matched or surpassed SOTA on standard benchmarks [89] | - | Transformer with domain adaptation |
| Multi-task DNN (SMILES) | - | 0.832 (STDNN), 0.840 (MTDNN) [88] | Pre-trained SMILES embeddings |
| Multi-task DNN (Fingerprint) | - | 0.824 (STDNN), 0.832 (MTDNN) [88] | Morgan fingerprint inputs |
BERT Architecture Implementation and Performance
BERT-based models have demonstrated particular effectiveness in molecular property prediction, especially in data-scarce scenarios common in drug discovery.
ChemLM employs a three-stage training process: (1) self-supervised pretraining on 10 million ZINC compounds using masked language modeling, (2) domain adaptation through further pretraining on task-specific unlabeled data with SMILES enumeration for augmentation, and (3) supervised fine-tuning for specific property prediction tasks [89]. This approach achieved substantially higher accuracy in identifying potent pathoblockers against Pseudomonas aeruginosa compared to state-of-the-art graph neural networks and language models, demonstrating its value for real-world drug discovery problems with limited training data [89].
Molecular BERT in Active Learning integrates pretrained BERT representations with Bayesian active learning, achieving equivalent toxic compound identification with 50% fewer iterations compared to conventional active learning [3]. The pretrained representations generate a structured embedding space that enables reliable uncertainty estimation despite limited labeled data, a critical advantage in low-data scenarios [3].
Figure 1: Three-stage training workflow for BERT-based molecular property prediction
Specialized Benchmarking Frameworks
Reproducible Tox21 Leaderboard
The recently introduced Tox21 leaderboard on Hugging Face addresses benchmark drift by re-establishing evaluation on the original challenge dataset and protocol [86]. The framework operates through automated API-based evaluation:
- Model Submission: Researchers submit models via Hugging Face Spaces with standardized FastAPI endpoints
- Inference: The leaderboard sends SMILES strings of the original 647 test compounds to the model's API
- Evaluation: Predictions are scored using the original challenge metric (mean AUC across 12 endpoints)
- Transparency: All baseline models and evaluation results are publicly accessible [86]
This infrastructure ensures historical fidelity while maintaining modern automation and transparency standards, providing a blueprint for other bioactivity prediction benchmarks suffering from similar drift issues [86].
MatBench for Materials Informatics
MatBench provides a standardized test suite of 13 supervised machine learning tasks for inorganic materials property prediction [43]. The benchmark includes a reference algorithm, Automatminer, which serves as a baseline for comparison. Automatminer automatically performs feature extraction using published materials featurizations, feature reduction, and model selection without user intervention [43].
Key findings from MatBench indicate that crystal graph neural networks tend to outperform traditional machine learning methods when approximately 10⁴ or more data points are available, while Automatminer achieved best performance on 8 of 13 tasks in the initial benchmark [43].
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Experimental Resources for Molecular Property Prediction
| Resource | Function | Application Examples |
|---|---|---|
| Tox21 Challenge Dataset | Benchmark for toxicity prediction models | 12,060 training + 647 test compounds with 12 toxicity endpoints [86] [87] |
| ClinTox Dataset | Clinical toxicity benchmarking | 1,484 compounds with FDA approval and clinical trial failure labels [3] [88] |
| MatBench Suite | Materials property prediction benchmark | 13 tasks with 312-132k samples across multiple property types [43] |
| Hugging Face Tox21 Leaderboard | Reproducible model evaluation | API-based automated testing on original Tox21 challenge set [86] |
| SMILES Embeddings | Molecular structure representation | Pre-trained embeddings capture structural relationships beyond fingerprints [88] |
| Scaffold Splitting | Evaluation strategy | Partitions data by core molecular structures to test generalization [3] |
| Automatminer | Automated materials ML pipeline | Reference algorithm for MatBench; performs automated featurization and model selection [43] |
Figure 2: Ecosystem of resources for molecular and materials property prediction research
Benchmarking on standardized datasets remains essential for tracking progress in molecular and materials property prediction. The evidence indicates that BERT-based architectures consistently achieve competitive performance across Tox21, ClinTox, and related benchmarks, with particular advantages in low-data scenarios through transfer learning and in active learning settings through improved uncertainty estimation.
Critical considerations for researchers include:
- The Tox21 benchmark drift issue necessitates careful interpretation of historical comparisons and preference for the reproducible leaderboard
- BERT-based models demonstrate strong performance with appropriate domain adaptation and pretraining
- Multi-task learning and advanced molecular representations (SMILES embeddings) generally outperform single-task models and traditional fingerprints
- Specialized benchmarking infrastructures like the Tox21 Leaderboard and MatBench provide essential standardization for meaningful comparison
Future directions should focus on extending these benchmarking principles to additional molecular property endpoints, improving model explainability in toxicity prediction, and further developing few-shot and zero-shot evaluation protocols for foundation models in chemistry and materials science.
The field of materials property prediction, particularly in drug discovery, has been revolutionized by deep learning architectures. Two dominant paradigms have emerged: Bidirectional Encoder Representations from Transformers (BERT), a transformer-based model excelling in sequential data processing, and Graph Neural Networks (GNNs), which specialize in relational data from graph structures. BERT leverages self-attention mechanisms to capture deep contextual relationships within sequential input data, making it powerful for understanding complex linguistic patterns or sequential molecular representations [90]. In contrast, GNNs operate on graph-structured data, iteratively aggregating information from neighboring nodes to capture topological relationships and structural patterns crucial for understanding molecular interactions and material properties [91]. This architectural divergence creates fundamental differences in their application strengths, performance characteristics, and suitability for specific research tasks in materials science and drug development.
Application Domains in Materials Property Prediction
BERT in Molecular Representation
BERT-based architectures have demonstrated remarkable success in molecular property prediction by treating chemical structures as sequential data. The Geometry-based BERT (GEO-BERT) framework incorporates three-dimensional molecular conformation data by introducing three distinct positional relationships: atom-atom, bond-bond, and atom-bond relationships [4]. This approach enables the model to capture spatial arrangements critical for predicting binding affinities and pharmacological properties. Similarly, pretrained BERT models have been integrated with Bayesian active learning to create data-efficient pipelines for molecular screening, achieving equivalent toxic compound identification with 50% fewer iterations compared to conventional active learning methods [9]. The sequential processing strength of BERT makes it particularly valuable for tasks involving molecular sequences, structural fingerprints, and textual data associated with materials.
GNNs in Structural Relationship Modeling
GNNs excel in directly modeling the inherent graph structure of molecules and materials, where atoms represent nodes and bonds represent edges. This native structural alignment enables GNNs to capture complex topological relationships and propagation patterns that sequential models might overlook [92]. In drug discovery, GNNs have been applied to predict drug-target interactions, protein function, and material properties by learning from the graphical representation of chemical structures [93]. The message-passing mechanism inherent in GNN architectures allows them to aggregate information from local neighborhoods in molecular graphs, making them particularly effective for predicting properties that emerge from localized structural motifs or intermolecular interactions.
Performance Comparison and Experimental Data
Quantitative Performance Metrics
Table 1: Performance comparison of BERT and GNN models across different tasks
| Model Architecture | Application Domain | Dataset | Key Metric | Performance | Reference |
|---|---|---|---|---|---|
| Dual-Stream BERT-GNN | Fake News Detection | FakeNewsNet | Accuracy | 99% | [92] |
| GEO-BERT | Molecular Property Prediction | DYRK1A Inhibitors | Novel Inhibitors Identified | 2 potent inhibitors (IC50: <1 μM) | [4] |
| BERT-DXLMA | Text Classification | Six Public Datasets | Overall Accuracy | Outperformed baseline methods | [90] |
| BERT with Bayesian Active Learning | Toxic Compound Identification | Tox21 & ClinTox | Data Efficiency | 50% fewer iterations needed | [9] |
| GNN Candidate-Job Matching | Recruitment Analytics | 8,360 candidates | Balanced Accuracy | 65.4% (GCN) vs 55.0% (MLP) | [94] |
Task-Specific Performance Analysis
The performance superiority of either BERT or GNN architectures heavily depends on the specific task and data characteristics. For textual analysis and sequential data processing, BERT-based models consistently achieve state-of-the-art performance. The BERT-DXLMA model, which integrates xLSTM architectures, demonstrated superior performance in text classification tasks across six public datasets, particularly in capturing deep semantic information and handling minority class samples in imbalanced data [90]. In contrast, GNNs show remarkable performance in relational tasks, as evidenced by a candidate-job matching system where GNN architectures achieved 65.4% balanced accuracy compared to 55.0% for multilayer perceptron baselines, correctly identifying 48.9% of qualified candidates versus only 8.5% for the MLP baseline [94].
Hybrid Architectures: Combining Strengths
Integrated BERT-GNN Frameworks
The most significant advances in materials property prediction have emerged from architectures that strategically combine BERT and GNN components. The Dual-Stream Graph-Augmented Transformer Model represents a pioneering approach that integrates BERT for deep textual representation with GNNs to model propagation structures of misinformation [92]. This architecture employs Graph Attention Networks (GAT) and Graph Transformers to extract contextual relationships while using an attention-based fusion mechanism to effectively integrate textual and graph embeddings for classification. Similarly, DynGraph-BERT creates a dynamic connection between BERT and GNN by exclusively using token embeddings to define and propagate graph structures, forcing BERT to redefine GNN graph topology to improve accuracy [91]. These hybrid approaches demonstrate that the combination of BERT's semantic understanding and GNN's structural reasoning capabilities can yield superior performance than either architecture alone.
Application in Drug Discovery
Hybrid BERT-GNN architectures have shown particular promise in drug discovery applications. A BERT-based Graph Neural Network was specifically designed for modeling drug-target binding affinity, leveraging BERT-style models pre-trained on vast quantities of both protein and drug data [93]. The encodings produced by each model are utilized as node representations for a graph convolutional neural network, modeling interactions without simultaneously fine-tuning both protein and drug BERT models. This approach significantly improved upon vanilla BERT baseline methods and former state-of-the-art methods on established drug-target interaction benchmarks, demonstrating the practical advantage of hybrid architectures in critical pathopharmacology applications.
Diagram 1: Hybrid BERT-GNN architecture showing dual-stream processing and fusion mechanism
Experimental Protocols and Methodologies
Benchmarking Standards
Robust evaluation of BERT and GNN models requires standardized experimental protocols across several dimensions:
Dataset Splitting: Scaffold splitting with 80:20 ratios is preferred for molecular datasets to create distinct training and testing sets that evaluate generalization capability. This method partitions molecular datasets according to core structural motifs identified by the Bemis-Murcko scaffold representation, ensuring train and test sets do not share identical scaffolds [9].
Evaluation Metrics: Comprehensive assessment includes accuracy, precision, recall, F1-score, and AUC-ROC for classification tasks. For regression tasks (e.g., binding affinity prediction), mean squared error and correlation coefficients are standard. Recent approaches also incorporate Expected Calibration Error measurements to assess uncertainty estimation reliability [9].
Baseline Comparisons: Properly optimized baselines are essential, including traditional machine learning (logistic regression, SVMs), single-modality deep learning models (CNNs, LSTMs), and established pre-trained models (BERT, RoBERTa) to ensure fair comparisons [95].
Implementation Details
Table 2: Key research reagents and computational tools for BERT-GNN experiments
| Research Reagent/Tool | Type | Function | Example Implementation |
|---|---|---|---|
| PyTorch | Deep Learning Framework | Model implementation and training | Used in Dual-Stream Graph-Augmented Transformer [92] |
| Hugging Face Transformers | Library | Pre-trained BERT models and utilities | Integration point for BERT components [92] |
| Graph Attention Networks (GAT) | GNN Architecture | Captures structural relationships with attention | Extracts contextual relationships in hybrid models [92] |
| Graph Transformers | GNN Architecture | Global attention on graph structures | Models propagation patterns in misinformation detection [92] |
| Bayesian Active Learning | Framework | Data-efficient model training | Reduces labeling iterations by 50% in toxicity screening [9] |
| Optuna | Hyperparameter Optimization | Automated hyperparameter tuning | Identifies optimal GNN architectures and embedding methods [96] |
Training Procedures
Training hybrid BERT-GNN models requires specialized procedures to handle the distinct characteristics of each component. The DynGraph-BERT approach incorporates dynamic graph construction, where BERT embeddings continuously redefine graph topology during training [91]. This method combines text augmentation with label propagation at test time, enhancing semi-supervised learning capabilities. For molecular property prediction, transfer learning approaches leverage BERT models pre-trained on large-scale molecular databases (e.g., 1.26 million compounds) before fine-tuning on specific property prediction tasks [9]. This strategy effectively disentangles representation learning from uncertainty estimation, leading to more reliable molecule selection in active learning scenarios.
Diagram 2: Experimental workflow for rigorous evaluation of BERT and GNN models
The performance comparison between BERT and GNN architectures reveals a complex landscape where architectural advantages are highly task-dependent. BERT-based models maintain superiority in tasks involving sequential data, deep semantic understanding, and transfer learning from large-scale pre-training. In contrast, GNNs excel in scenarios requiring explicit modeling of structural relationships, topological patterns, and propagation dynamics. For materials property prediction and drug discovery applications, hybrid architectures that leverage the strengths of both paradigms demonstrate the most promising results, achieving state-of-the-art performance across multiple benchmarks.
Future research directions should focus on developing more efficient fusion mechanisms, enhancing model interpretability for scientific discovery, and adapting these architectures for low-data scenarios common in novel material development. As the field progresses, the integration of BERT and GNN methodologies will likely become increasingly seamless, potentially giving rise to unified architectures that dynamically adapt their processing strategy based on data characteristics and task requirements. For researchers and drug development professionals, this evolving landscape offers powerful tools for accelerating material discovery and optimization, provided they carefully match architectural strengths to their specific predictive challenges.
The application of machine learning to materials property prediction represents a frontier where modern deep learning architectures and traditional algorithms compete for dominance. Within this domain, researchers and drug development professionals must navigate a complex landscape of algorithmic options, primarily divided between transformer-based architectures like BERT and classical machine learning approaches such as Random Forests (RF) and Gaussian Processes (GP). Each paradigm offers distinct advantages: BERT leverages pre-trained representations on vast molecular datasets, Random Forests provide robust performance on structured tabular data, and Gaussian Processes deliver principled uncertainty quantification essential for scientific discovery. This guide objectively compares these approaches within materials property prediction contexts, examining their theoretical foundations, empirical performance, implementation requirements, and suitability for different research scenarios in materials science and drug development.
Performance Comparison: Quantitative Metrics Across Domains
Empirical Performance in Materials Property Prediction
Table 1: Comparative performance of ML algorithms across material property prediction tasks
| Algorithm | Application Domain | Key Performance Metrics | Uncertainty Quantification | Data Efficiency |
|---|---|---|---|---|
| BERT-based Models | Polymer property prediction [97] | 1st place in NeurIPS Open Polymer Prediction Challenge 2025 (over 2,240 teams) | Limited native capability | Requires pretraining on large datasets (e.g., 1.26M compounds [24] [9]) |
| Random Forests | Environmental mapping [98] | Spatial RF variants outperform standard RF over short prediction distances | Limited to ensemble variance | Works well with small to medium datasets |
| Gaussian Processes | Thermophysical properties [99] | R² ≥0.85 for 5/6 properties, ≥0.90 for 4/6 properties tested | Native, well-calibrated uncertainty estimates | Struggles with very large datasets |
| Deep Gaussian Processes | HEA property prediction [100] | Best performance for correlated material properties with heteroscedastic data | Captures both epistemic and aleatoric uncertainty | Medium data requirements |
Computational Requirements and Scalability
Table 2: Computational characteristics and implementation requirements
| Algorithm | Training Speed | Inference Speed | Scalability | Hyperparameter Complexity |
|---|---|---|---|---|
| BERT-based Models | Slow (requires pretraining) | Medium | High memory requirements (24GB GPU for large molecules [97]) | High (Optuna-tuned learning rates, batch sizes [97]) |
| Random Forests | Fast | Fast | Excellent for tabular data [101] | Low to medium |
| Gaussian Processes | Medium | Slow for large datasets | O(n³) complexity limits dataset size | Medium (kernel selection crucial [99]) |
| Deep Gaussian Processes | Slow | Medium | Better than standard GPs for complex data [100] | High |
Experimental Protocols and Methodologies
BERT Implementation for Molecular Property Prediction
The winning solution in the NeurIPS Open Polymer Prediction Challenge 2025 exemplifies a sophisticated BERT implementation pipeline for property prediction [97]. The methodology employs a multi-stage approach:
Data Preparation and Augmentation: SMILES representations of polymers are converted to canonical form with deduplication. Data augmentation generates 10 non-canonical SMILES per molecule using
Chem.MolToSmiles(..., canonical=False, doRandom=True, isomericSmiles=True), expanding training data tenfold.Two-Stage Pretraining:
- Stage 1: An ensemble of BERT, Uni-Mol, AutoGluon, and D-MPNN models generates property predictions for 50,000 polymers from the PI1M dataset.
- Stage 2: BERT models undergo pretraining on a pairwise comparison classification task, predicting which polymer exhibits higher/lower property values across all five target properties, excluding pairs with similar values.
Fine-Tuning Protocol: Implementation uses AdamW optimizer with no frozen layers, one-cycle learning rate schedule with linear annealing, automatic mixed precision, and gradient norm clipping at 1.0. The backbone learning rate is set one order of magnitude lower than the regression head to prevent overfitting.
Inference: Generates 50 predictions per SMILES with median aggregation for final prediction. This approach demonstrates that general-purpose BERT (ModernBERT-base) outperformed chemistry-specific models like ChemBERTa and polyBERT in polymer property prediction [97].
Random Forest Implementation for Spatial Prediction
Spatial Random Forest implementations for environmental mapping follow a structured protocol [98]:
Spatial Feature Engineering: Incorporate spatial coordinates and relationships as features, including:
- Geographical coordinates and spatial lag variables
- Distance-based features to relevant landmarks or boundaries
- Leave-one-out Ordinary Kriging predictions based on out-of-bag errors as spatial covariates
Spatial Variant Implementation: Six spatial RF variants are benchmarked against universal kriging and multiple linear regression, with RF-OOB-OK (using ordinary kriging predictions based on out-of-bag error) emerging as a consistently well-performing method.
Validation Strategy: Employ spatial cross-validation techniques that account for geographical autocorrelation, assessing performance over different prediction distances to evaluate how well spatial structure is captured.
Hyperparameter Tuning: Optimize tree depth, number of trees, and minimum sample leaf size through random search or Bayesian optimization, though RF is generally robust to hyperparameter settings [102].
Gaussian Process Implementation for Uncertainty-Aware Prediction
Gaussian Process protocols emphasize uncertainty quantification alongside point predictions [99] [103]:
Kernel Selection: Test multiple covariance functions (Radial Basis Function, Matérn, Rational Quadratic) to capture different smoothness assumptions in the data, with the GCGP method proving robust to variations in kernel choice [99].
Mean Function Specification: For hybrid GCGP models, integrate Group Contribution method predictions as mean functions, enabling the GP to learn and correct systematic biases in baseline predictions.
Hyperparameter Optimization: Maximize marginal likelihood to optimize kernel hyperparameters (length scales, variance) rather than cross-validation, providing a principled Bayesian approach to parameter tuning.
Sparse Approximation: For larger datasets, implement sparse GP variants using inducing points to maintain computational tractability while preserving uncertainty quantification.
Diagram 1: Comparative workflow architectures of BERT, Gaussian Process, and Random Forest pipelines for materials property prediction.
Table 3: Key computational tools and resources for materials informatics
| Tool/Resource | Function | Application Context |
|---|---|---|
| ModernBERT-base | General-purpose foundation model | Polymer property prediction; outperformed chemistry-specific BERT variants [97] |
| AutoGluon | Automated tabular model training | Feature engineering and model selection for Random Forests and gradient boosting [97] |
| GPy/GPyTorch | Gaussian Process implementation | Flexible kernel specification and uncertainty quantification [99] [100] |
| Uni-Mol-2-84M | 3D molecular representation | Captures spatial molecular structure for property prediction [97] |
| RDKit | Molecular descriptor generation | Computes 2D/3D molecular features for traditional ML inputs [97] |
| Optuna | Hyperparameter optimization framework | Tunes learning rates, batch sizes, and architecture decisions [97] |
| Chem.MolToSmiles | SMILES augmentation | Generates non-canonical SMILES for data expansion in BERT training [97] |
Integration Patterns and Hybrid Approaches
Bayesian Active Learning with Pretrained Representations
Recent advances demonstrate the power of integrating pretrained BERT representations with Bayesian active learning frameworks for drug design [24] [9]. This hybrid approach effectively disentangles representation learning from uncertainty estimation, addressing a critical limitation in low-data scenarios:
Representation Learning: MolBERT, pretrained on 1.26 million compounds, provides high-quality molecular embeddings that structure the chemical space meaningfully before fine-tuning on specific property prediction tasks.
Uncertainty Estimation: Bayesian acquisition functions like BALD (Bayesian Active Learning by Disagreement) and EPIG (Expected Predictive Information Gain) leverage these representations to select informative molecules for labeling, achieving equivalent toxic compound identification with 50% fewer iterations compared to conventional active learning [9].
Experimental Design Formalization: The framework treats molecule selection as a Bayesian experimental design problem, maximizing expected information gain about model parameters through strategic compound selection.
Deep Gaussian Processes with Architectural Priors
For complex multi-task prediction scenarios with correlated material properties, Deep Gaussian Processes (DGPs) infused with machine learning-based priors have demonstrated superior performance [100]:
Hierarchical Architecture: DGPs stack multiple GP layers to capture complex, non-stationary patterns in high-entropy alloy property data that conventional GPs cannot represent effectively.
Prior Integration: Machine learning-derived priors guide the DGP to better capture inter-property correlations and input-dependent uncertainty in hybrid datasets combining experimental and computational properties.
Heteroscedastic Modeling: The hierarchical structure naturally handles varying noise levels across different measurement types and conditions common in materials informatics.
Diagram 2: Hybrid Bayesian active learning workflow combining pretrained BERT representations with Bayesian experimental design for efficient drug discovery.
The comparative analysis reveals that algorithm selection in materials property prediction depends critically on research constraints and objectives. BERT-based approaches excel when abundant pretraining data exists and representation quality dominates other considerations, particularly in molecular design applications. Random Forests remain formidable for tabular datasets with strong feature representations, offering robust performance with minimal hyperparameter tuning. Gaussian Processes provide unparalleled uncertainty quantification essential for experimental design and safety-critical applications, though at computational cost for large datasets. Hybrid approaches that combine pretrained representations with Bayesian methods represent the cutting edge, enabling data-efficient active learning that accelerates materials discovery and drug development. Researchers should prioritize BERT for representation-heavy tasks with transfer learning potential, Random Forests for rapid prototyping on structured data, and Gaussian Processes when uncertainty quantification is paramount for decision-making.
This guide provides an objective comparison of performance metrics for various BERT-based architectures in molecular property prediction, a critical task in modern drug discovery. The evaluation focuses on Mean Absolute Error (MAE) and accuracy within the broader context of data efficiency, offering researchers a clear framework for model selection.
In computational drug discovery, molecular property prediction serves as a cornerstone for identifying promising therapeutic candidates. The shift from traditional machine learning to sophisticated BERT-based architectures has created a need for nuanced performance evaluation. While accuracy is commonly used for classification tasks, Mean Absolute Error (MAE) provides a straightforward, interpretable measure of average error magnitude for regression problems, calculated as the average of absolute differences between predicted and actual values [104]. As labeled molecular data is often scarce and expensive to obtain, data efficiency—the ability of a model to achieve high performance with limited training examples—has emerged as a critical metric for evaluating model practicality [9].
Comparative Analysis of BERT Architectures
The table below compares key BERT-based architectures on performance metrics relevant to molecular property prediction.
Table 1: Performance Comparison of BERT-based Architectures for Molecular Property Prediction
| Model Architecture | Key Features | Reported Performance | Data Efficiency | Best Use Cases |
|---|---|---|---|---|
| BERT with Bayesian Active Learning [9] | Pretrained BERT + Bayesian experimental design | Achieved equivalent toxic compound identification with 50% fewer iterations vs. conventional AL | High | Drug toxicity prediction with limited labeled data |
| Geometry-based BERT (GEO-BERT) [4] | Incorporates 3D molecular conformations | Identified two novel DYRK1A inhibitors (IC50: <1 μM); optimal performance on multiple benchmarks | Information missing | Predicting properties dependent on 3D molecular structure |
| Self-Conformation-Aware Graph Transformer (SCAGE) [105] | Multitask pretraining on ~5M compounds | Significant improvements across 9 molecular properties and 30 structure-activity cliff benchmarks | Information missing | General molecular property prediction with interpretability needs |
| Domain-Adapted Transformers [106] | Further-trained on domain-specific data & objectives | Performance plateau at 400-800K pre-training molecules; significant gains from domain adaptation | Medium | ADME property prediction with limited domain data |
Experimental Protocols and Methodologies
Protocol 1: BERT with Bayesian Active Learning
This methodology focuses on optimizing the data acquisition process itself [9].
- Objective: To maximize predictive performance while minimizing experimental labeling costs.
- Dataset Preparation: Publicly available molecular datasets (e.g., Tox21, ClinTox) are split into training and test sets using scaffold splitting to ensure generalization to novel molecular structures. An initial small, balanced set of labeled molecules (e.g., 100) is selected, with the remainder treated as an unlabeled pool [9].
- Model Components: A BERT model (e.g., MolBERT) pretrained on large-scale unlabeled molecular databases (e.g., 1.26 million compounds) provides high-quality molecular representations [9].
- Active Learning Loop: The core iterative process involves:
- The model is trained on the current set of labeled molecules.
- Acquisition Function: A Bayesian acquisition function (e.g., BALD - Bayesian Active Learning by Disagreement) calculates the informativeness of every molecule in the unlabeled pool. BALD selects points where the model's parameters are most uncertain, maximizing information gain [9].
- The top-ranked molecules are selected for "labeling" (simulating a costly experiment) and added to the training set.
- Performance Measurement: Model accuracy (e.g., in toxic compound identification) is tracked against the number of active learning iterations or the total number of labeled molecules used [9].
Protocol 2: Geometry-Integrated BERT (GEO-BERT) Evaluation
This protocol evaluates models that incorporate spatial molecular information [4].
- Objective: To assess the impact of 3D structural information on prediction accuracy.
- Data Preparation: Molecular structures are converted to their low-energy 3D conformations using force field methods (e.g., MMFF). The 3D coordinates are integrated into the model input [4] [105].
- Model Training: The geometry-aware model (e.g., GEO-BERT) is trained to predict specific molecular properties (e.g., binding affinity). A baseline model without 3D information is trained for comparison [4].
- Performance Validation: Model performance is quantified using MAE or accuracy on benchmark datasets. Crucially, prospective validation is performed: top-ranked predictions are tested in wet-lab experiments to determine actual inhibitory concentration (IC50) [4].
Protocol 3: Domain Adaptation for Transformers
This protocol measures gains from tailoring a generic model to a specific chemical domain [106].
- Objective: To improve performance on specific molecular property endpoints (e.g., ADME: Absorption, Distribution, Metabolism, Excretion).
- Model Setup: A transformer model is first pre-trained on a large, general-purpose molecular dataset (e.g., GuacaMol).
- Domain Adaptation: The model is further trained ("adapted") on a smaller, domain-relevant unlabeled dataset (e.g., molecules relevant to solubility). This step often uses chemically informed objectives like Multi-Task Regression (MTR) of physicochemical properties [106].
- Evaluation: The domain-adapted model is fine-tuned and evaluated on downstream ADME tasks. Its performance is compared against the base model and other benchmarks using MAE or ROC-AUC [106].
Figure 1: Workflow of three primary BERT-based approaches for molecular property prediction, highlighting their paths to optimizing key performance metrics.
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Computational Tools and Datasets
| Tool/Resource | Type | Primary Function in Research |
|---|---|---|
| Tox21 & ClinTox Datasets [9] | Data | Benchmark datasets for training and evaluating models on toxicity prediction tasks. |
| MolBERT / Pretrained BERT [9] [71] | Software | Provides foundational molecular representations, transferring knowledge from large-scale unlabeled data. |
| Bayesian Acquisition Functions (BALD, EPIG) [9] | Algorithm | Quantifies model uncertainty to identify the most informative molecules for labeling in active learning cycles. |
| Molecular Conformation Generators (MMFF) [105] | Software | Computes stable 3D structures of molecules from their 2D representations, essential for geometry-aware models. |
| SMILES/DeepSMILES Strings [71] | Data | Standardized string-based representations of molecular structure used as input for sequence-based models like BERT. |
| Domain-Specific ADME Datasets [106] | Data | Curated data for properties like solubility and permeability, used for domain adaptation to improve model specificity. |
Discussion and Strategic Recommendations
The comparative analysis reveals that no single BERT architecture universally outperforms others across all metrics. The optimal choice is heavily dependent on the specific research context and constraints.
For projects with severely limited labeled data or where labeling is prohibitively expensive, the BERT with Bayesian Active Learning framework is the most strategic choice. Its proven ability to reduce data requirements by up to 50% while maintaining performance offers a significant practical advantage [9]. When predicting properties known to be highly dependent on 3D molecular geometry (e.g., protein-ligand binding), GEO-BERT and similar geometry-integrated models are preferable, as their incorporation of spatial information leads to higher accuracy and successful experimental validation [4]. For well-defined tasks where a moderate amount of labeled data exists and the chemical domain is narrow (e.g., ADME prediction), Domain-Adapted Transformers provide a balanced approach, leveraging chemically informed objectives to achieve robust performance without ultra-large-scale pre-training [106].
In conclusion, researchers must prioritize their constraints—whether data, computational resources, or the nature of the target property—to select the model architecture that best optimizes the trade-offs between MAE, accuracy, and data efficiency. Future work will likely focus on hybrid models that integrate the strengths of these approaches, such as combining 3D awareness with efficient active learning paradigms.
In the field of computational drug discovery, the ability to predict the properties and interactions of novel, previously uncharacterized chemical compounds is a fundamental challenge. Zero-shot learning (ZSL) has emerged as a powerful paradigm to address this, enabling models to make accurate predictions for compounds they have never encountered during training [107]. This capability is particularly vital within BERT-based architecture research for materials property prediction, as it directly tests a model's capacity to generalize beyond its training data and leverage learned fundamental principles of chemistry and structural relationships.
This guide provides a comparative analysis of state-of-the-art ZSL methodologies, focusing on their application to unseen compounds. It details experimental protocols, presents quantitative performance data, and outlines the essential toolkit for researchers working at the intersection of BERT architectures and molecular property prediction.
Comparative Analysis of Zero-Shot Learning Methodologies
Table 1: Comparison of Zero-Shot Learning Approaches for Compound and Target Interaction
| Method / Model | Core Approach | Application Context | Key Performance Metrics (Unseen Compounds/Targets) |
|---|---|---|---|
| PSRP-CPI [108] | Protein subsequence reordering pretraining & length-variable augmentation. | Compound-Protein Interaction (CPI) prediction. | Improved baseline performance in Unseen-Compound, Unseen-Protein, and Unseen-Both scenarios. |
| ZeroBind [109] | Protein-specific meta-learning & subgraph information bottleneck. | Drug-Target Interaction (DTI) prediction. | AUROC: 0.8139 (Inductive set with unseen proteins/drugs). |
| Mol-BERT / MolRoPE-BERT [63] | Exploration of positional embeddings (absolute, rotary, etc.) in BERT. | Molecular property prediction from SMILES/DeepSMILES. | Increased accuracy and generalization in zero-shot molecular property prediction. |
| Simulation-Driven GZSL [110] | Semantic mapping from simulated single-fault to compound-fault data. | Bearing compound fault diagnosis (Conceptual parallel to molecular systems). | High-accuracy classification of unseen compound fault classes in a Generalized ZSL setting. |
| Fine-tuned BERT (BioGottBERT) [111] | Task-specific fine-tuning of a specialized BERT model. | Symptom extraction from clinical text (Non-molecular, but a ZSL benchmark). | F1 score: 0.84 (Outperformed general-purpose zero-shot models like GLiNER and Mistral). |
Detailed Experimental Protocols
PSRP-CPI for Compound-Protein Interaction
The PSRP-CPI method addresses the challenge of modeling complex interdependencies between protein subsequences that are critical for binding but are often non-adjacent in the sequence [108].
- Objective: To predict interactions between compounds and proteins, including in zero-shot scenarios involving unseen compounds or proteins.
- Procedure:
- Pretraining: A protein encoder, typically a multi-layer Transformer, is pretrained using a subsequence reordering pretext task. Protein sequences are divided into segments, randomly shuffled, and the model is trained to predict the correct original order. This forces the model to learn structural and functional dependencies between non-adjacent subsequences.
- Data Augmentation: A length-variable protein augmentation strategy is applied by processing sequences of varying lengths. This enhances model robustness and performance, especially when training data is limited.
- Fine-tuning: The pretrained encoder is integrated with baseline CPI models (e.g., GraphDTA, HyperattentionDTI) and fine-tuned on specific CPI benchmark datasets. The model is then evaluated on test sets split into "Seen-Both," "Unseen-Compound," "Unseen-Protein," and "Unseen-Both" categories.
- Evaluation: Performance is measured using standard metrics like Area Under the Receiver Operating Characteristic curve (AUROC) and Area Under the Precision-Recall Curve (AUPRC) across the different zero-shot scenarios [108].
ZeroBind for Drug-Target Interaction
ZeroBind tackles the generalization problem for unseen proteins and drugs through a meta-learning framework that trains protein-specific models [109].
- Objective: To predict drug-target interactions in both zero-shot and few-shot settings for novel proteins and drugs.
- Procedure:
- Task Formulation: The problem is framed as a meta-learning task, where training a model for a single protein's DTI prediction is considered one "task."
- Network-Based Negative Sampling: To address data imbalance, non-interacting drug-target pairs are generated by pairing drugs and proteins from different network communities, creating robust negative examples.
- Model Architecture:
- Graph Encoder: A Graph Convolutional Network (GCN) learns embeddings from the molecular graph of the drug and the graph structure of the protein.
- Subgraph Information Bottleneck (SIB): This module identifies maximally informative and compressive subgraphs within the protein graph as potential binding pockets, enhancing interpretability and performance.
- Task Adaptive Self-Attention: A module that learns the importance of different protein-specific tasks for the meta-learner.
- Meta-Training: The model is trained using the MAML++ framework. The meta-learner is optimized over many protein-specific tasks so it can quickly adapt to new proteins.
- Evaluation: Model performance is rigorously assessed on independent transductive, semi-inductive, and fully inductive test sets using AUROC and AUPRC, with the inductive set representing the true zero-shot scenario for unseen proteins and drugs [109].
BERT with Positional Embeddings for Molecular Property Prediction
This approach investigates how different positional encoding strategies in BERT architectures affect the model's understanding of molecular structure from SMILES strings, thereby influencing zero-shot generalization [63].
- Objective: To increase the accuracy and generalization of molecular property prediction by exploring various positional embeddings (PEs) within a transformer-based framework.
- Procedure:
- Pretraining: A BERT model is pretrained on a large corpus of unlabeled molecular SMILES or DeepSMILES strings (e.g., ~7.9 million instances) using Masked Language Modeling (MLM). Different types of PEs, including absolute, relativekey, relativekey_query, and sinusoidal, are experimentally compared.
- Fine-tuning: The best-performing pretrained model for each PE type is subsequently fine-tuned on downstream tasks involving labeled data for specific molecular properties. This includes datasets related to COVID-19, bioassay data, and other biological properties.
- Zero-Shot Evaluation: The model's proficiency is assessed by predicting properties for molecular representations not seen during pretraining or fine-tuning, demonstrating its ability to generalize to novel regions of chemical space [63].
The Scientist's Toolkit: Essential Research Reagents & Materials
Table 2: Key Reagents and Computational Tools for ZSL Research
| Item / Resource | Function / Description | Relevance to ZSL Experiments |
|---|---|---|
| SMILES / DeepSMILES Strings [63] | Text-based representations of molecular structures. | The primary input data for BERT-based models pretrained on chemical structures. |
| BindingDB, CHEMBL, PDBbind [109] | Public databases of drug-target binding data and compound information. | Source of curated, labeled data for training and fine-tuning DTI/CPI models. |
| Graph Neural Networks (GNNs) [109] | Neural networks that operate directly on graph-structured data. | Used to learn embeddings from the native graph structure of molecules and proteins. |
| Meta-Learning Algorithms (e.g., MAML++) [109] | Frameworks for training models on a distribution of tasks to enable fast adaptation. | Core to methods like ZeroBind for generalizing to proteins/drugs with no or few samples. |
| Dynamic Model Simulation Data [110] | Simulated data generated from physical/engineering models of system behavior. | Used as auxiliary information to train semantic mapping models when real labeled data for "unseen" classes is scarce. |
| Subgraph Information Bottleneck (SIB) [109] | A technique to extract maximally informative subgraphs from a larger graph. | Identifies critical functional units (e.g., binding pockets in proteins), improving model interpretability and focus. |
The evaluation of model generalization to unseen compounds is a critical frontier in AI-driven drug discovery. As the comparative analysis shows, methods like PSRP-CPI and ZeroBind demonstrate that through innovative pretraining tasks, meta-learning frameworks, and sophisticated model architectures, robust zero-shot prediction is achievable. Furthermore, the exploration of positional embeddings in BERT models underscores the importance of fundamental architectural choices in understanding molecular syntax. These approaches, supported by the detailed experimental protocols and tools outlined herein, provide researchers with a powerful arsenal to advance the state of the art in predicting the properties and interactions of the vast expanse of unexplored chemical space.
Conclusion
The integration of BERT architecture into materials and molecular property prediction marks a significant paradigm shift, moving beyond traditional feature engineering and graph-based models. The key takeaways underscore BERT's superior ability to handle data scarcity through innovative pretraining and multitask learning, its flexibility enabled by cross-modal transfer and architectural hybrids, and its proven performance against state-of-the-art methods on critical benchmarks. For biomedical and clinical research, these advances promise a faster, more cost-effective path to drug candidate screening and optimization. Future directions will likely involve training on even larger, multimodal datasets, deeper integration with generative models for molecular design, and a stronger focus on model interpretability to build trust and provide actionable insights for scientists, ultimately accelerating the journey from novel compound discovery to clinical application.
References
- 1. Why Materials Discovery Is Key to the Net Zero Transition [https://netzeroinsights.com/resources/materials-discovery/]
- 2. Accelerating materials innovation with AI [https://www.innovationnewsnetwork.com/accelerating-materials-innovation-with-ai/62755/]
- 3. Molecular property prediction using pretrained-BERT and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12020163/]
- 4. Geometry-based BERT: An experimentally validated deep ... [https://www.sciencedirect.com/science/article/pii/S2095177925002825]
- 5. Enhancing composition-based materials property ... [https://arxiv.org/html/2511.03371v1]
- 6. Transformer-generated atomic embeddings to enhance ... [https://www.nature.com/articles/s41467-025-56481-x]
- 7. Going beyond SMILES enumeration for data augmentation ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12409607/]
- 8. In silico methods for drug-target interaction prediction - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC12570315/]
- 9. Molecular property prediction using pretrained-BERT and ... [https://jcheminf.biomedcentral.com/articles/10.1186/s13321-025-00986-6]
- 10. Hybridization of SMILES and chemical-environment-aware ... [https://www.nature.com/articles/s41598-025-01890-7]
- 11. SMILES alignment: a dynamic programming approach for the ... [https://bmcbioinformatics.biomedcentral.com/articles/10.1186/s12859-025-06278-y]
- 12. Stacked variational autoencoder with SMILES-based drug ... [https://www.sciencedirect.com/science/article/abs/pii/S0925231225025093]
- 13. Drug Discovery SMILES-to-Pharmacokinetics Diffusion ... [https://arxiv.org/html/2408.07636v2]
- 14. AI in Chemistry — Identifying Structural Similarities through ... [https://medium.com/@aakash.ezhilan/ai-in-chemistry-identifying-structural-similarities-through-smile-notation-5a88651bf343]
- 15. BERT vs GPT vs LLaMA: AI Model Comparison 2025 [https://www.byteplus.com/en/topic/560409]
- 16. DrugBERT: a BERT-based approach integrating LDA topic ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12326809/]
- 17. Transformer Architecture Explained With Self-Attention ... [https://www.codecademy.com/article/transformer-architecture-self-attention-mechanism]
- 18. An Interpretable Soft-Sensor Framework for Dissertation Peer ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12568271/]
- 19. Comparison of BERT implementations for natural language ... [https://www.sciencedirect.com/science/article/pii/S2352914822002763]
- 20. Analyzing Sleep Behavior Using BERT-BiLSTM and Fine ... [https://medinform.jmir.org/2025/1/e70753]
- 21. A comparative study of pre-trained language models for ... [https://bmcmedinformdecismak.biomedcentral.com/articles/10.1186/s12911-022-01967-7]
- 22. What are masked language models? [https://www.ibm.com/think/topics/masked-language-model]
- 23. Masked language modeling [https://huggingface.co/docs/transformers/main/tasks/masked_language_modeling]
- 24. a data-efficient approach to drug design [https://pubmed.ncbi.nlm.nih.gov/40270038/]
- 25. [2406.06553] Ensemble Model With Bert,Roberta and Xlnet ... [https://arxiv.org/abs/2406.06553]
- 26. A systematic review of deep learning chemical language ... [https://jcheminf.biomedcentral.com/articles/10.1186/s13321-024-00916-y]
- 27. LLM-enhanced Multi-level Knowledge Distillation for ... [https://www.sciencedirect.com/science/article/abs/pii/S0950705125012365]
- 28. A Comparative Review of Large Language Models in ... [https://www.mdpi.com/2227-9717/13/9/2680]
- 29. Benchmarking Pretrained Molecular Embedding Models ... [https://arxiv.org/html/2508.06199v3]
- 30. A framework for evaluating the chemical knowledge and ... [https://www.nature.com/articles/s41557-025-01815-x]
- 31. Assessing the Chemical Intelligence of Large Language ... [https://arxiv.org/html/2505.07735v1]
- 32. Rethinking chemical research in the age of large language ... [https://www.nature.com/articles/s43588-025-00811-y]
- 33. A roadmap for large language models in chemical research [https://engineering.cmu.edu/news-events/news/2025/06/24-roadmap-for-llms.html]
- 34. Recent advances in molecular representation methods and ... [https://www.nature.com/articles/s44386-025-00017-2]
- 35. Pre-trained molecular language models with random ... [https://www.nature.com/articles/s44387-025-00029-3]
- 36. Two-Stage Pretraining for Molecular Property Prediction in ... [https://arxiv.org/html/2411.03537v1]
- 37. A quantum-transformer hybrid architecture for polymer ... [https://www.sciencedirect.com/science/article/pii/S0927025625002939?dgcid=rss_sd_all]
- 38. A review on multi-task metric learning | Big Data Analytics [https://bdataanalytics.biomedcentral.com/articles/10.1186/s41044-018-0029-9]
- 39. Multi-task learning [https://en.wikipedia.org/wiki/Multi-task_learning]
- 40. Randomized SMILES strings improve the quality of molecular ... [https://jcheminf.biomedcentral.com/articles/10.1186/s13321-019-0393-0]
- 41. Optimizing Evaluation Metrics for Multi-Task Learning via ... [https://arxiv.org/abs/2210.05935]
- 42. Invalid SMILES are beneficial rather than detrimental to ... [https://www.nature.com/articles/s42256-024-00821-x]
- 43. the Matbench test set and Automatminer reference algorithm [https://www.nature.com/articles/s41524-020-00406-3]
- 44. LLM4Mat-Bench: Benchmarking Large Language Models ... [https://arxiv.org/abs/2411.00177]
- 45. Cross-modality material embedding loss for transferring ... [https://www.nature.com/articles/s41524-025-01723-1]
- 46. A large-scale human toxicogenomics resource for drug- ... [https://www.nature.com/articles/s41467-025-65690-3]
- 47. Multimodal deep learning for chemical toxicity prediction ... [https://www.nature.com/articles/s41598-025-95720-5]
- 48. VitroBert: modeling DILI by pretraining BERT on in vitro data [https://jcheminf.biomedcentral.com/articles/10.1186/s13321-025-01048-7]
- 49. A Bayesian active learning platform for scalable ... [https://www.nature.com/articles/s41467-024-55287-7]
- 50. A Bayesian active learning platform for scalable combination ... [https://pubmed.ncbi.nlm.nih.gov/39746987/]
- 51. Comparative Study of Molecular Descriptors and AI-Based ... [https://pubmed.ncbi.nlm.nih.gov/41235418/]
- 52. AIVIVE: a novel AI framework for enhanced in vitro to in ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12469192/]
- 53. TOXRIC: a comprehensive database of toxicological data and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9825425/]
- 54. Feature-Assisted Machine Learning for Predicting Band ... [https://www.mdpi.com/2079-4991/14/5/445]
- 55. Predicting band gap from chemical composition [https://arxiv.org/html/2501.02932v1]
- 56. A Survey of AI for Materials Science: Foundation Models, LLM Agents... [https://arxiv.org/html/2506.20743v1]
- 57. Accurate prediction of band gap of two-dimensional ... [https://www.sciencedirect.com/science/article/abs/pii/S2542529325001300]
- 58. Accurate Prediction of Experimental Band Gaps from Large ... [https://arxiv.org/abs/2311.13778]
- 59. A general-purpose material data extraction pipeline from... property [https://www.nature.com/articles/s41524-023-01003-w?error=cookies_not_supported&code=0a9504d9-8e24-4111-a123-50543863a498]
- 60. Positional embeddings and zero-shot learning using BERT for... [https://link.springer.com/article/10.1186/s13321-025-00959-9]
- 61. Positional Embeddings in Transformer Models [https://iclr-blogposts.github.io/2025/blog/positional-embedding/]
- 62. Complete Summary of Absolute, Relative and Rotary Position ... [https://azizbelaweid.substack.com/p/complete-summary-of-absolute-relative]
- 63. Positional embeddings and zero-shot learning using BERT for ... [https://jcheminf.biomedcentral.com/articles/10.1186/s13321-025-00959-9]
- 64. Moving closer to experimental level materials property ... [https://www.nature.com/articles/s41598-022-15816-0]
- 65. A BERT model generates diagnostically relevant semantic ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9053264/]
- 66. Enhancing materials property prediction by leveraging ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC6874674/]
- 67. QFFN-BERT: An Empirical Study of Depth, Performance ... [https://arxiv.org/html/2507.02364v1]
- 68. A quantum-transformer hybrid architecture for polymer ... [https://www.sciencedirect.com/science/article/abs/pii/S0927025625002939]
- 69. Quantum-Embedded Graph Neural Network Architecture ... [https://pubmed.ncbi.nlm.nih.gov/40785363/]
- 70. Hybrid quantum neural networks show strongly reduced ... [https://www.nature.com/articles/s41598-025-88177-z]
- 71. Positional embeddings and zero-shot learning using BERT for ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11800558/]
- 72. A topic-aware classifier based on a hybrid quantum- ... [https://link.springer.com/article/10.1007/s00521-023-08706-7]
- 73. Quantum intelligence in drug discovery: Advancing insights ... [https://www.sciencedirect.com/science/article/abs/pii/S135964462500176X]
- 74. The Future of Drug Discovery: 2025 as the Inflection Year ... [https://modelmedicines.com/newsroom/the-future-of-drug-discovery-2025-as-the-inflection-year-for-hybrid-ai-and-quantum-computing]
- 75. Explainable Artificial Intelligence: A Perspective on Drug ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12472777/]
- 76. Explainable AI: Visualizing Attention in Transformers - Comet [https://www.comet.com/site/blog/explainable-ai-for-transformers/]
- 77. jessevig/bertviz: BertViz: Visualize in NLP Models ( Attention ...) BERT [https://github.com/jessevig/bertviz]
- 78. Deconstructing BERT , Part 2: Visualizing the Inner... - KDnuggets [https://www.kdnuggets.com/2019/03/deconstructing-bert-part-2-visualizing-inner-workings-attention.html]
- 79. SMG- BERT : integrating stereoscopic information and chemical... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10348360/]
- 80. Ensemble Model With Bert , Roberta and Xlnet For Molecular ... property [https://arxiv.org/html/2406.06553v1]
- 81. From SMILES Tokens to Molecular Embeddings (2/3) [https://medium.com/@anrizal05/chemical-foundation-models-from-smiles-tokens-to-molecular-embeddings-2-3-a118e7589686]
- 82. Improving the quality of chemical language model outcomes ... [https://jcheminf.biomedcentral.com/articles/10.1186/s13321-023-00725-9]
- 83. Comparing SMILES and SELFIES tokenization for ... [https://www.nature.com/articles/s41598-024-76440-8]
- 84. Hybridization of SMILES and chemical-environment-aware ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12081657/]
- 85. Hybrid fragment-SMILES tokenization for ADMET prediction in ... [https://bmcbioinformatics.biomedcentral.com/articles/10.1186/s12859-024-05861-z]
- 86. A Reproducible Leaderboard for the Tox21 Challenge [https://arxiv.org/html/2511.14744]
- 87. Tox21 Machine Learning Data Set [https://bioinf.jku.at/research/DeepTox/tox21.html]
- 88. Accurate clinical toxicity prediction using multi-task deep ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10039880/]
- 89. Domain adaptable language modeling of chemical ... [https://www.nature.com/articles/s42004-025-01484-4]
- 90. BERT-DXLMA: Enhanced representation learning and ... [https://www.sciencedirect.com/science/article/abs/pii/S0925231224020964]
- 91. DynGraph-BERT: Combining BERT and GNN Using ... [https://www.mdpi.com/2227-9709/12/1/20]
- 92. Dual stream graph augmented transformer model ... [https://www.nature.com/articles/s41598-025-05586-w]
- 93. Modelling Drug-Target Binding Affinity using a BERT ... [https://pubmed.ncbi.nlm.nih.gov/34892183/]
- 94. Graph Neural Networks for Candidate-Job Matching [https://link.springer.com/article/10.1007/s41019-025-00293-y]
- 95. Are We Really Making Much Progress in Text Classification ... [https://arxiv.org/html/2204.03954v6]
- 96. A powerful fusion of language and graphs [https://www.sciencedirect.com/science/article/abs/pii/S0925231225012457]
- 97. Winning Recipe For Predicting Polymer Properties [https://xebia.com/blog/winning-recipe-for-predicting-polymer-properties/]
- 98. An assessment of spatial random forests for environmental ... [https://www.sciencedirect.com/science/article/pii/S136481522500310X]
- 99. Enhanced thermophysical property with uncertainty... prediction [https://pubs.rsc.org/en/content/articlelanding/2025/me/d5me00126a]
- 100. [2506.14828] Accurate and Uncertainty-Aware Multi-Task Prediction ... [https://arxiv.org/abs/2506.14828]
- 101. Random Forests in 2025 — Why This “Old” Algorithm Still ... [https://medium.com/data-science-collective/random-forests-in-2025-why-this-old-algorithm-still-wins-on-real-world-data-after-20-years-142b25183453]
- 102. Comparison of methods for tuning machine learning model hyper-parameters: with application to predicting high-need high-cost health care users | BMC Medical Research Methodology | Full Text [https://bmcmedresmethodol.biomedcentral.com/articles/10.1186/s12874-025-02561-x]
- 103. Combining Deep Learning and Gaussian for Processes ... Accurate [https://scisimple.com/en/articles/2025-07-13-combining-deep-learning-and-gaussian-processes-for-accurate-predictions--a3wnon8]
- 104. Is This Old Accuracy Metric Still Surprisingly Relevant? [https://covidstatus.dps.illinois.edu/is-still-reasonably-accurate]
- 105. A self-conformation-aware pre-training framework for ... [https://www.nature.com/articles/s41467-025-59634-0]
- 106. Transformers for Molecular Property Prediction: Domain ... [https://arxiv.org/html/2503.03360v3]
- 107. Zero-Shot and Few-Shot Learning: Learning with Limited Data [https://deepfa.ir/en/blog/zero-shot-few-shot-learning-limited-data]
- 108. Zero-Shot Learning with Subsequence Reordering ... [https://arxiv.org/html/2507.20925v1]
- 109. ZeroBind: a protein-specific zero-shot predictor with ... [https://www.nature.com/articles/s41467-023-43597-1]
- 110. Simulation-data Driven Generalized Zero-Shot Learning for ... [https://www.sciencedirect.com/science/article/abs/pii/S0950705125006410]
- 111. Optimized BERT-based NLP outperforms Zero-Shot ... [https://www.medrxiv.org/content/10.1101/2025.04.21.25326037v1]